Abstract
E-26 Transformation Specific (ETS) transcription factors are known to cause various cancers and their aberrant expression has been related to various oncogenic processes like metastasis and angiogenesis. Currently very few drugs exist which can directly target oncogenic transcription factor and none of them are approved for human use. To design ETS transcription factor targeting small molecules, an understanding of their regulation in cancers is essential. In this review, we have discussed upstream transcriptional regulations of ETS factors that includes epigenetic alterations in DNA like methylation and acetylation, alternate promoter, fusion gene formation, promoter affinity/specificity, recruitment of cofactors/repressors, etc. Additionally, we have also detailed the regulatory post translational modifications like phosphorylation, acetylation and ubiqutinylation for various ETS family members. These transcriptional and translational alterations regulate the expression of all ETS family members and can provide rational for drug designing in future.
Keywords
ETS, Transcription, Phosphorylation, Acetylation, Ubiqutinylation, Regulation
Introduction- The ETS Family History
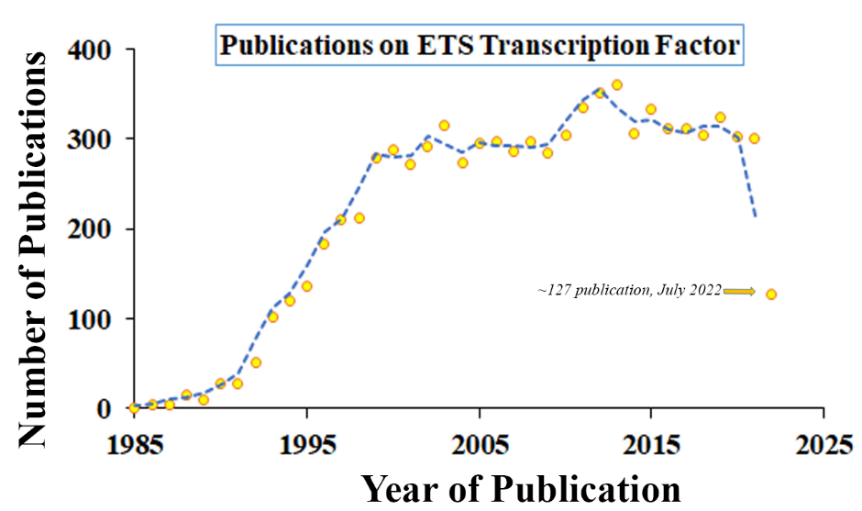
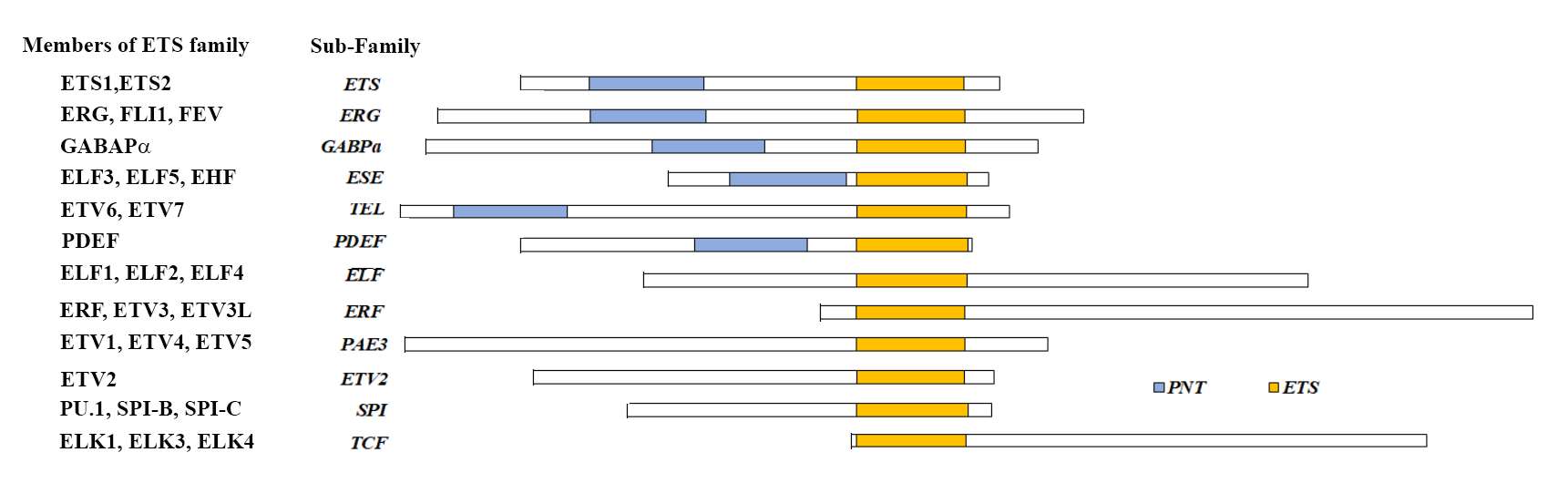
ETS1; the first E-Twenty-six Transformation Specific transcription factor member capable of cellular transformation; was first discovered and characterized in the early eighties by several groups [1-3]. Since then, more than 7,500 scientific articles have made their entries in National Library of Medicine (PubMed search) for the term ‘ETS Transcription Factor’, indicating the immense importance of these factors in biological sciences and diseases (Figure 1). These metazoan specific proteins consists of 12 subfamilies sharing a conserved 85 amino acid long DNA binding domain called the ‘ETS domain’ which recognizes a consensus purine rich GGA(A/T) core [4,5].Till date based on conserved sequence homology and functions, 28 ETS family members in humans (27 in mice) were identified (Figure 2) [6,7]. ETS family have various physiological roles like hematopoiesis, immune cell maturation, neuronal and vascular development which were prominent in knock out mouse models (Table 1) [6]. However, in context of cancers most ETS family members are considered to be oncogenic capable of transformation. Overexpression of important ETS factors like ERG, FLI1, ETS1, ETV1, ETV4/5/6, ELK, SPi1, are directly involved in initiation of leukemias, prostate cancer, breast cancer, sarcomas, astrocytoma, hepatic carcinoma, melanomas, etc. [8-20]. Additionally, these ETS factors also regulate various oncogenic processes like angiogenesis, invasiveness, epithelial mesenchymal transition (EMT), stemness, metabolic reprogramming, immune evasiveness, deregulation of checkpoints, etc. by activation or repression of their target genes [14,17,19,21-27]. In this review, we will focus on regulation of ETS transcription factors in cancers.
Figure 1. Publications on ETS transcription factors with respect to timeline.
Figure 2: ETS family members with conserved domains.
|
ETS Factor |
Phenotype/physiological role in knock-out mice |
Refs |
|
ETS1 |
T cell survival, activation and maturation defects, Defects in B cell auto immune response, NK cells lineage defects, vascular inflammation and remodeling defects, endothelial cell survival defects, congenital and developmental defects of kidney. |
[28-34] |
|
ETS2 |
Extra embryonic membrane defects, endothelial cell survival defects, Thymocyte maturation defects. |
[32,35-37] |
|
ERG |
Defects in angiogenesis |
[38] |
|
FLI1 |
Disruption of hematopoietic lineage |
[16] |
|
ETV1 |
Arterial remodeling and Arrhythmia |
[39] |
|
ETV 2 |
Lack of hematopoietic and hematoendothelial lineage; Defects in vasculogenesis |
[40-44] |
|
ETV 4 |
Estrogen signaling in endometrium |
[45] |
|
ETV 5 |
Defects in spermatogenesis; breakdown of cellular homeostasis in alveolar type II(AT2) cells |
[46,47] |
|
ELK 1 |
Development of male sterility |
[48] |
|
ELK 4 |
Defects in thymocytes |
[49] |
|
ELK 3 |
Increased synthesis of nitric oxide (Reactive nitrogen species); delayed post-natal retinal angiogenesis in mice |
[50,51] |
|
GABPα |
Encouraged invasion and metastasis in papillary and follicular thyroid carcinoma |
[52,53] |
|
ELF 1 |
Reduced transcriptional responses to cytokine interferon-β |
[54] |
|
ELF 4 |
Causes autoinflammatory and immunodeficiency diseases |
[55] |
|
ETV 6 |
Defects in embryonic hematopoiesis |
[56] |
|
Spi B |
Inhibition of plasmacytoid dendritic cell development |
[57] |
|
Spi C |
Defective inflammatory response and iron metabolism in macrophage |
[58] |
|
Spi 1 |
Retardation of erythropoiesis |
[59] |
|
Erf |
Maintained embryonic stemness, highly expressed pluripotency |
[60] |
|
Elf 3 |
Severe alteration of tissue architecture in small intestine, poor villus formation, abnormal morphogenesis, terminal differentiation of absorptive enterocytes |
[61]
|
|
Elf 5 |
Absence of alveologenesis |
[62] |
|
SPDEF |
Lack of conjunctival goblet cells and formation of dry eye |
[63] |
Family Hierarchy: Transcriptional Regulation of ETS Factors in Cancers
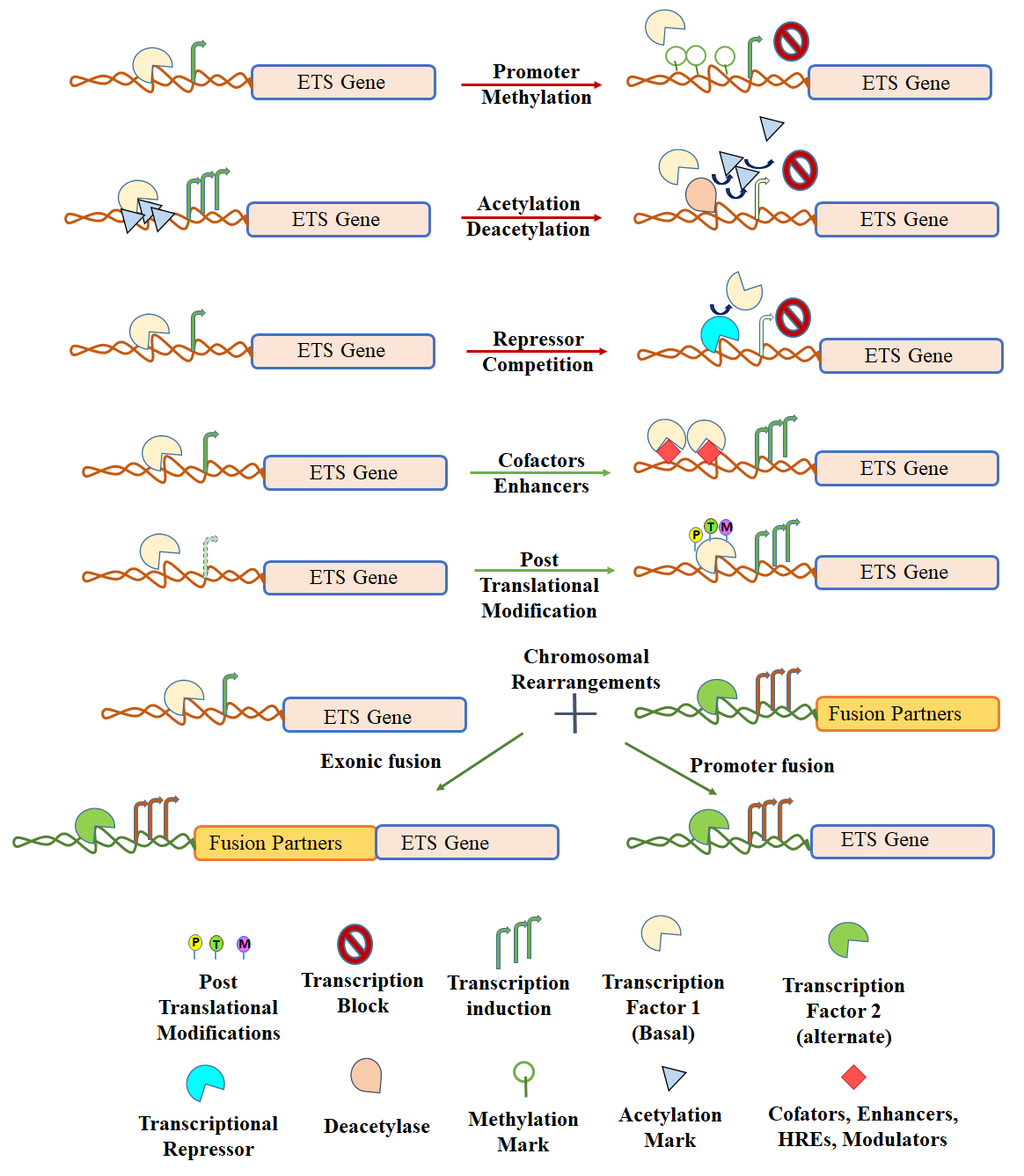
All gene expressions are regulated at their promoters by various transcription factors, DNA modifications, cofactors and enhancers. Deregulation of transcription factor binding at promoters often triggers cancer initiation [64-67]. Regulation of ETS family members is modulated by various mechanisms of transcriptional regulation like promoter methylation/acetylation, promoter replacement by gene fusion formation, activation/repression of upstream cofactors/transcription factors, alternate promoter activation/specificity; which are represented in Figure 3. Since ETS family consist of a large number of genes, we will focus on a few important transcriptional regulatory mechanisms of ETS members that are involved in human cancers.
Figure 3: Mechanisms of regulation: ETS factors.
ETS family members; ETS1 & ETS2
The TATA box less ets1 gene, is regulated positively at its promoter by upstream regulatory factors like AP1, AP2, HIF2alpha, and ETS1 itself [68-70]. Other growth factors like, TNF-alpha, HGF can indirectly induces ETS1 mRNA in cancers [71,72]. Interestingly, Retinoic acid was shown to alter ets1 expression in both positive and negative manner with context of cell milieu [73-75]. Recently, Nf-KB1 and Nfatc2 was shown to bind ETS1 promoter resulting in its up regulation and promotion of breast cancer invasiveness [76]. As a negative regulator, tumor suppressor p53 was shown to repress ETS1 expression [77]. In parallel, ets2 gene is transcriptionally repressed by v-Myb and ERF in various cancers [78,79].
ERG family members; ERG, FLI1, FEV
ERG is known as a primary driver in many cancers. Interestingly, 30 transcript variants of ERG have been identified due to multiple promoters and their epigenetic regulation via methylation [80-82]. Surprisingly, ERG can form gene-fusion with other gene promoter or regulatory elements like TMPRSS2, EWS, TLS/FUS etc. resulting in deregulation of ERG fusion transcripts in prostate cancer, Ewing sarcoma, leukemia [15,50,83-90]. Occasionally, ERG fusions can drive wild type ERG’s transcription [91]. Similarly oncogenic FLI1 can form fusions with other genes like EWS, FUS, etc resulting in aberrant expression of FLI1 in cancers like Ewing sarcoma [92]. Wild type FLI1 promoter is induced by ETS1, ETS2, ELF1 or FLI1 itself in a GATA factor dependent manner, but is inhibited by ETS factor Tel/ETV6 [93-95]. FLI1 is regulated by ETV2 in a feed forward loop where both ETV2 and FLI1 can upregulate FLI1 transcriptionally [96]. ETS factor coding the FEV (Fifth Ewing Variant) gene is regulated by transcriptional activation of GATA sites at its promoter [97]. Interestingly, FEV is implicated as a tumor suppressor and is down regulated in prostate cancer [98]. FEV translocation (2;16) with either FUS or ETV1 generates a high expressing FUS-FEV or FEV-ETV1 fusion gene in <10% of Ewing sarcoma [99,100].
ESE family members; ELF3, ELF5
ELF3 (ETS transcription factor 3) is transcriptionally induced in trophoblasts by EVT-specific super-enhancers [101]. In lung adenocarcinoma, ELF3 locus is often amplified and hypomethylated resulting in increased expression [102]. While high ELF3 expression promotes epithelial mesenchymal transitions in liver cancer and breast cancer, the role is reversed in case of ovarian cancer [103-107]. ELF5 (ETS transcription factor 5) expression in embryonic stem cells and breast epithelial cells are regulated by promoter methylation resulting in lineage determination and mammary gland development respectively [108-110]. ELF5 owing to multiple promoter have tissue specific expression which are often deregulated in cancers [108,111]. ELF5 promotes basal phenotype in breast cancer by inhibiting ERalpha and Foxa1 [112,113]. Additionally, ELF5 was shown to inhibit breast cancer metastasis and EMT by transcriptionally repressing factors like SNAIL2, CD24 indicating a diabolic behavior in oncogenesis [114,115].
TEL family members; ETV6
ETV6 or TEL (Translocation ETS Leukemia) is often deregulated in leukemia, colorectal and breast cancer [79,116-118]. Deficiency in ETV6 expression often results in thrombocytopenia and patients are often predisposed to leukemia [119]. Promoter methylation or regulation via other factors like AKIRIN1, COMMD9, DYRK4, JUNB, and SRP72 have been implicated in regulating ETV6 expression in metabolism and cancers [120,121]. Recently, RBPJ protein was shown to transcriptionally regulate ETV6 and its fusion in glioblastoma [122]. ETV6 can form more than 30 types of gene fusions in various cancers [123,124]. Factors regulating wildtype ETV6 promoter might also regulate ETV6 fusion genes like ETV6-NTRK3, ETV6-RUNX1 and ETV6-JAK2 in various cancers [122,125-128].
ELF family members; ELF1, ELF2
ELF1 (E74-like factor 1), associated with erythroid maturation can be regulated at its promoter by transcription factor PU.1 [129]. Rb gene was reported to interact with ELF1 inhibiting T cell activation by down regulation of ELF1 transcription program [130]. ELF1 acts as tumor suppressor in prostate cancer; as deletion of ELF1 results in senescence and increased docetaxel susceptibility [131]. On the contrary, ELF1 promotes glioma development by activating GFI1/FBW7 axis via MEIS1 factor indicating cancer specific roles [132]. Not much is known about regulation of ELF2 in cancers except that it is involved in processes like angiogenesis and invasiveness [133,134].
PEA3 members; ETV1, ETV4, ETV5
Homologues of ETV1/4/5 forms the PEA3 family which are involved in various cancers [135]. Although ETV1 (ETS variant 1) is mainly regulated by androgen, other factors like postnatal maturation gene signaling also contribute to regulation of its expression [21,136,137]. ETV1 can regulate various oncogenic properties in cancers of prostate, colorectal, breast, etc [12,137-140]. Additionally, ETV1 gene can form fusions with inducible promoter of TMPRSS2, OR51E2-ETV1, EWSR1-ETV1 and regulates various cancer [90,141,142]. Whereas ETV4 can regulate its own transcript expression in a tissue dependent manner in various cancers [143-146]. ETV4 can also form fusions like TMPRSS2-ETV4, SLC45A3-ETV4, DDX5-ETV4, FUS/ETV4, NCOA2-ETV4 in various cancer [99,141,147-149]. Similarly, ETV5 expression is regulated by ALK in a MAPK dependent manner in neuroblastoma [150]. Promoter methylation of ETV5 also upregulates it’s expression in colon cancer [151]. ETV5 promotes oncogenic processes like myometrial infiltration in endometrial cancer, proliferation and anchorage independent growth, cell adhesion in ovarian cancer [152-154]. ETV5 also forms fusion in prostate cancer with partners like TMPRSS2 and SLC45A3 [155].
ELK family member; ELK1, ELK3 and ELK4
ELK1(ETS Like-1) expression is regulated by factors like ERK kinase in association with TCF-SRF (serum response factor) complexes in astrocytomas, androgens in bladder cancer [156,157]. High ELK expression is associated with cancers of colon, urinogenital and thyroid. ELK3 expression is up regulated by ribosomal kinase RSK2 or Ras/ERK pathways promoting cancers [158-160]. Hypoxia can downregulate ELK3 expression [161,162]. ELK4 among the members can form a SLC45A3-ELK4 fusion in prostate cancer regulated by androgen signaling [163]. ELK4 also promotes gastric cancer and glioblastoma through anti-apoptotic mechanism [164,165].
Family Business: Post Translational Modifications of ETS Protein Products in Cancer
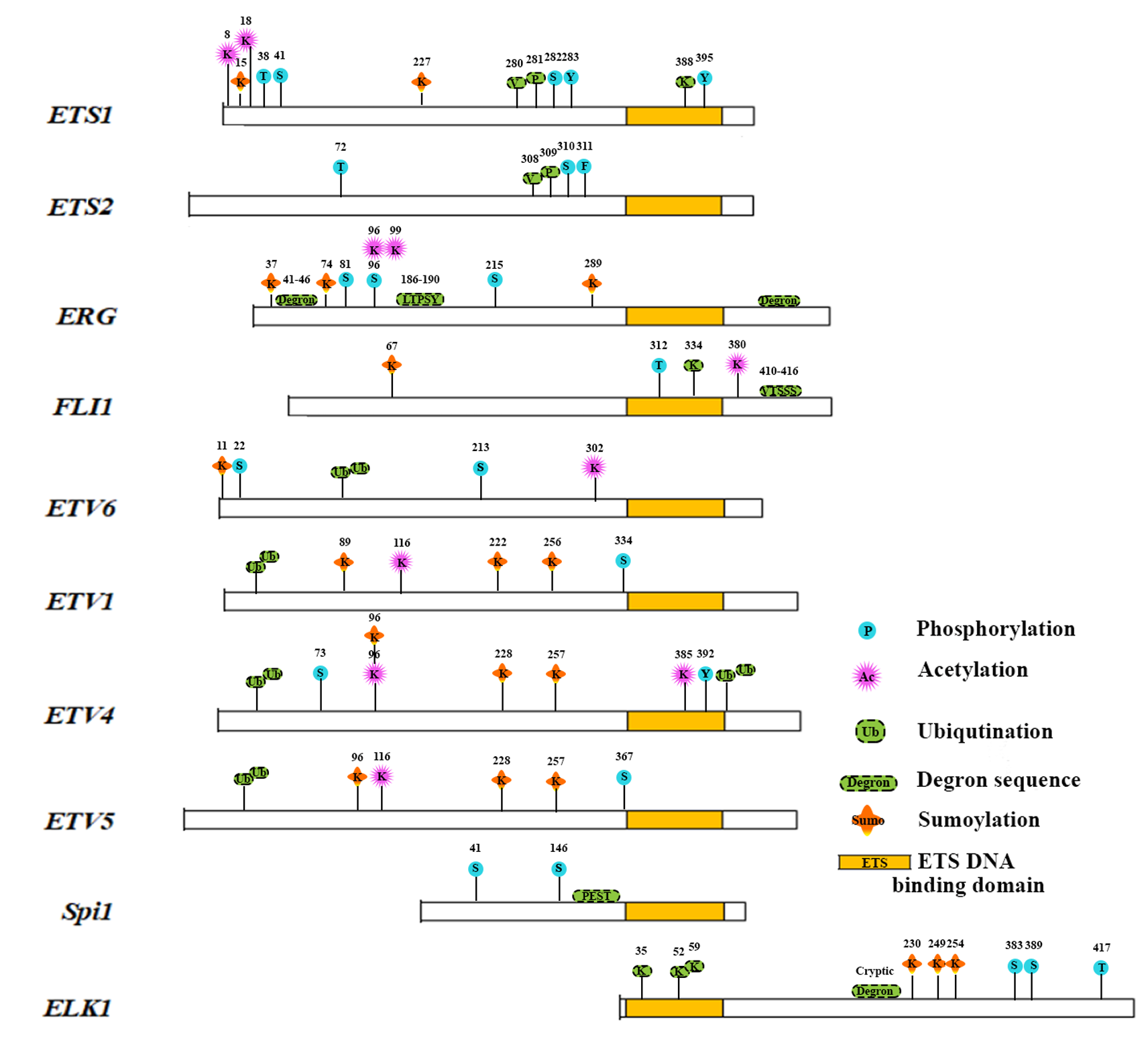
Apart from ETS-DNA binding domain, which is conserved among all family members of ETS transcription factors, there are other important domains among the various members. For example, Pointed domain (PNT), which is present among ETS, ERG, GABPα, ESE and TEL sub-families is responsible for homo-oligomerization, hetero-oligomerization and transcriptional repression [166-168]. Other domains, includes the TAD (Transactivation domain) domain responsible for cofactor docking, activation/repression of transcriptional functions of ETS family members, B-box SRF interacting region responsible for enhanced DNA binding are found among members of ETS family [169-173]. Post translational modifications (PTMs) like phosphorylation, acetylation, ubiqutinylation, glycosylation etc., in protein domains are often associated with structure and function based regulation of transcription factors including ETS family in cancers [174]. Various PTMs of ETS transcription factor families represented across domains and families in Figure 4 and their effect on function and regulation of these factors in cancer will be discussed below.
Figure 4: Post Translational Modifications of ETS family proteins.
Phosphorylation: The bread winner
Various growth factors, stress, mitogenic signals activate serine/threonine kinase MAPK/ERK; which are known to phosphorylate various ETS family members and initiate their functions in cancers. For example; transcription factor ERG was shown to be consecutively phosphorylated by ERK kinase at Serine 215 (S215) and S96 residue resulting in loss of EZH2-Suz12 polycomb repressor complex and activation of ERG mediated metastatic program in prostate cancer [175]. Recently, AKT kinase directed modulation of ERK -ERG phosphorylation axis was identified; where presence of active AKT dictates luminal fates of prostate cancer cells instead of EMT program [176,177]. Meanwhile MAPK/ERK2 kinase can phosphorylate ERG oncoprotein; resulting in hematopoietic stem and progenitor cells (HSPCs) proliferation and activation of oncogenic programs in acute myeloid leukemia (AML) and T-acute lymphoblastic leukemia (T-ALL) [178]. Although other members of the subfamily; FLI1 and FEV were shown to be in vitro phosphorylated by MAPK/ERK, there is no direct role of ERK phosphorylation on regulating FLI1 or FEV mediated oncogenesis [179]. MAPK/ERK kinases not only phosphorylates ETS subfamily members ETS1 and ETS2 in various cancers but are also transcriptionally regulated by ETS1/2 for maintenance of cancer program [180-182]. ETS1 phosphorylation at Threonine38 (T38) and S41 via ERK results in CBP/p300 mediated transcriptional activation, increased invasiveness and EMT in breast cancer [180,183-186]. While ETS2 phosphorylation by ERK1/2 were prominent in tamoxifin resistance breast tumors [187]. Multiple phosphorylation by MAPK/ERK cascade are also reported in other family members like i) Phosphorylation at S383,S389, T417 residues of ELK1 oncoprotein resulting in enhanced transactivation, proliferation, anti-apoptotic mechanisms and metabolic reprogramming in diverse cancers [173,188-192] ii) Phosphorylation at S73 residue of ETV4 (Pea3 subfamily) oncoprotein resulting in stabilization of ETV4 via blockage of ubiquitinylation signals in colorectal cancers [193]. While phosphorylation of another Pea3 family member; ETV1 oncoprotein in the activation domain (90-160 amino acid) resulted in activation of ETV1 transcriptional program in various cancers [194,195].
Other than mitogen activated kinases, depending upon cell milieu; various kinases affects ETS factor phosphorylation and subsequent function/regulation in cancers. For example; i) calcium signaling modulators CAMP kinases act as inhibitory signals, as CAMKII mediated phosphorylation inhibited transactivation of both ETS1 and ETS2 family members via multiple phosphorylation adjacent to DNA binding domain [196-199]. Interestingly, crosstalk between multiple phosphorylation are needed for complete inhibitory effects [196,197]. ii) AGC kinase family members like PKA, PKC kinases have been reported in phosphorylation of ETS factors. Both ETS1 and ETS2 proteins are stabilized and activated by PKC kinase in various cancers [200,201]. While ERG subfamily member FLI1 was phosphorylated at Threonine 312 (T312) by PKC delta kinase in a c-abl dependent manner resulting in FLI1 activation, stem cell differentiation and immune modulation [202-204]. PKC alpha mediated phosphorylation of Pea3 sub family members ETV1 and ETV5 at S334 and S367 respectively altering their DNA binding specificity towards high/low affinity promoters results in tumorigenic signaling [205,206]. iii) Src kinase mediated phosphorylation of ETS1/ETS2 at tyrosine 283 (Y283)/ Phenylalanine 311 (F311) has been implicated in various cancers [207]. Src mediated phosphorylation prevents binding and subsequent ubiqutinylation of ETS factors by COP1 E3 ligase [207,208]. iv) SPI sub family member Spi-1/PU.1 was reported to be phosphorylated by AKT kinase at S41 resulting in transactivation [209,210]. While Spi-B was reported to be phosphorylated by Casein Kinase II at multiple sites (S37, S129, S144 and S146) resulting in increased protein stability and decreased transactivation in cancers [211].
Acetylation: The frugal son
Acetylation of transcription factors including ETS family members often dictates subcellular distribution, DNA affinity, stability, transcriptional activity and results in cancers [212-215]. The pioneer member ETS1 is acetylated and dissociated by p300 (histone acetyl transferase; HAT) from chromatin during activation of miR-192 [216,217]. Again, ETS1 can be acetylated by CBP (HAT) at lysine 8 (K8) and K18 within the N-terminal domain, facilitating interaction with BRD4 and activating VEGF mediated angiogenesis program [218]. ERG is acetylated by p300 at K96/K99 residues promoting BRD4 regulated hematopoietic transcriptional program in acute myeloid Leukemia and invasiveness in case of prostate cancer [219,220]. Other ERG subfamily member FLI1 was reported to be acetylated at lysine 380 by p300/CBP-associated factor (PCAF) resulting in decreased FLI1 stability and DNA binding affinity [221]. While HDAC1 was identified as a deacetylase for FLI1 increasing DNA binding and transcriptional program associated with FLI1 directed oncogenesis [222]. On the contrary, EWS-FLI1 chimeric oncoprotein, is acetylated by both p300 and PCAF at the C-terminal FLI1 domain (at K240, K252, and K380 lysine number corresponds to wildtype FLI1) resulting in stronger DNA binding and transcriptional activation of FLI1 [223]. Pea3 family members ETV1 and ETV5 are acetylated by p300/PCAF at K116 residue resulting in transactivation and stability of the oncoproteins [114,224,225]. While Pea3 member ETV4 exhibits multiple p300 mediated lysine acetylation promoting transactivation functions [226,227]. Other acetylation include, HAT Tip-60 mediated acetylation and co-repression of ETV6/TEL that mediates transcriptional programs [228] and p300 mediated acetylation and subsequent ubiquitinylation of ELF5 in breast cancer [229].
Ubiquitylation & sumoylation: The prodigal son
Protein turnover is often regulated via proteasomal pathway via PTMs like ubiquitylation and sumoylation. Ubiquitin moiety can bind to lysine(K) residues within proteins creating mono (K63) or poly(K48) ubiquitin chains, which may then act as signals for proteasomal or other pathways [230-232]. The turnover of ETS factors are tightly regulated by crosstalk between PTMs resulting in ubiqutination mediated degradation. For example, COP1 E3 ubiquitine ligase has been reported to ubiqutinylate various ETS factors at specific valine(V)/proline(P) residue pairs followed by phosphorylation events. ETS1 and ETS2 are poly-ubiquitinylated at V280/P281 and V308/P309 residues following serine phosphorylation resulting in their proteasomal degradation by E3 ubiquitin ligase COP1 [207,231,233]. In fact ETS1 and ETS2 are protected from this ubiquitinylation by consecutive phosphorylation at their adjacent sites S282/Y283 and S310/Y311 by CAMKII and SRC kinases [231]. PEA3 family members ETV1/ETV4/ETV5 are also subjected to poly-ubiqutinylation at V/P sites and subsequent proteasomal degradation by COP1 [234,235]. Interestingly, COP1 is suggested to act as a tumor suppressor and prognostic marker and its deregulation has been reported in many cancer [235-239]. ERG family proteins lacking COP1 binding sites are resistant to COP1 mediated ubiqutinylation [235]. However, E3 ubiqutine ligase SPOP (Speckle-type POZ protein) has been reported to degrade ERG by ubiqutinylation of specific degron motifs at N-terminus(41-46 amino acid) and C-terminus (23-27 a.a) of ERG resulting in suppression of ERG positive prostate cancer progression [240,241]. ERG can be degraded by SCF E3 ligase complex where a typical phospho degron (186LTPSY190) is recognized by the ligase for ubiquitination [242,243]. ERG Family members; FLI1 is also ubiqutinylated at K334 in its wild type form and at residue K380 when it exists as the EWS-FLI1 fusion oncoprotein [244]. Recently it is observed that, SPOP E3 ligase can regulate both FLI1 and its fusion protein’s turnover by binding to a VTSSS” degron motif of EWS-FLI1/FLI1 (462-468aa/410-416aa), which is a prerequisite for polyubiqutinylation and subsequent proteasomal degradation [245]. While TEL family member ETV6 is ubiquitinylated at SAM domain by F-box family ligase FBXL6 resulting in proteasomal degradation [246,247]. ELK1 can be a substrate for both poly and mono ubiqutinylated [248]. ELK1 is reversibly mono-ubiqutinylated at site K35, K52, K59 resulting in impaired DNA binding activity [249]. While F box ubiquitine ligase, FBXO25 can poly ubiqutinylate ELK1 at a cryptic degron motif (amino acids 167-196) resulting in its degradation [250,251].
SUMO (small ubiquitin-like modifier) based modification facilitated by the SUMO conjugation enzyme system results in sumoylation of protein leading to various signaling events. With a few exceptions, most ETS factors undergo transcriptional repression, degradation or loss of functionality upon sumoylation. For example, sumoylation of ETS1 at K15 and K227 residues by E2 SUMO-conjugating enzyme Ubc9 and a E3 SUMO ligase, PIASy results in defective transactivation without any effect on turnover of the protein [233]. In acute myeloid leukemia; ERG is hyper sumoylated at K37, K74, and K289 by PIAS4 SUMO E3 ligase resulting in enhanced stability of ERG protein [252]. On the contrast, ERG family member FLI1 fails to transactivation when sumoylated by sumo E3 ligase PIASxα/ARIP3 at K67 site [253]. All Pea3 family member ETV1/4/5 are known to be sumoylated [135]. These ETS factors are sumoylated at conserved lysine residues (K89/96, K222/228, and K256/257) resulting in their altered transcriptional efficiency [254,255]. Tel family member ETV6 is also sumoylated at K11 residue by PIAS3 ligase resulting in repression of ETV6 transcriptional activity [256]. Interestingly, UBC9, ubiquitin conjugating enzyme has been shown to promote ETV6 sumoylation resulting in localization to cell-cycle-specific nuclear speckles in leukemias housing a TEL/AML fusion [257]. ELK1’s nucleus to cytoplasmic shuttling is modulated by sumoylation at K230, K249, or K254. Recently, ELK1 transactivation was connected to p38MAPK mediated phosphorylation, which promotes sumoylation and downregulation of activity [258]. PTMs like Ubiquitination and sumoylation in transcription factors are strictly regulated and as with others, deregulation of these PTMs results in enhanced activity of ETS family in various cancer [259-261].
Few other types of PTMs like methylation and glycosylation are also reported for a few ETS factors. In prostate cancer, TMPRSS2-ERG fusion oncogenic activity is induced by EZH2 mediated methylation of the oncoprotein at K362 residue [262]. Activation of the Ets transcription factor Elf-1 requires multiple glycosylation in patients with lupus disease [263].
Conclusion, Be Careful of that Family
Owing to their implications in cancers, ETS family of transcription factors are important drug targets at present [264,265]. However, ETS factors also have physiological roles, thus off target effects of drugs should be considered during drug selection [266-268]. Interestingly, ETS factors like ERG, ETV1, FLI1, ETV6, etc., can form gene fusions which acts as drivers of many cancers [269,270]. These fusions are active drug targets and various inhibitors/drugs against these fusion ETS factors are listed in Table 2. A few studies on ETS regulation based on cellular milieu also indicated unknown factors that might play vital role during chemotherapy. For example, glutamine starvation can reduce expression of ETS1 and alter its translocation, thereby reducing its transcriptional activity in ovarian cancers [271]. Alternatively, in a hormone dependent cancer like prostate, androgens rich environment provides induction for TMPRSS2-ERG [272]. This can be advantageous during chemotherapy, as it is reported that phosphorylation of ERG fusions during DNA damage results in its degradation; independent of androgen response [242]. Recently, metabolic coactivator PGC1-alpha was reported to induce ERG fusions transactivation during metabolic stress in cancers indicating a metabolic vulnerability during resistant cancer development [273]. Thus, modern day cancer therapy targeting ETS transcription factors must take into account the cellular milieu and stress response during drug designing.
|
Drugs |
ETS fusion target |
Cancer |
Mechanisms |
Ref. |
|
TK-216 |
EWS-FLI 1 |
Ewing sarcoma |
Decrease oncogene expression and increase tumor suppressor genes, induce apoptosis NCT02657005 |
[274]
|
|
Englerin A |
EWS-FLI 1 |
Ewing sarcoma |
Induction of cell cycle arrest and cell death; Inhibition of cell proliferation; reduction of EWS-FLI 1 phosphorylation and DNA binding |
[264] |
|
Midostaurin (PKC412) |
EWS-FLI 1 |
Ewing sarcoma |
Inhibited cell growth and promoted apoptosis |
|
|
Mithramycin |
EWS-FLI1 |
Ewing sarcoma |
Causes repression of H3 acetylation and association with EWS-FLI1 protein thereby reduction in oncogenesis |
[275] |
|
Trabectedin |
EWS-FLI1 |
Ewing sarcoma |
Inhibits the expression of EWS-FLI1 and tumor associated macrophage |
|
|
Trabectedin + Oliparib |
EWS-FLI1 |
Ewing sarcoma |
Inhibits the expression of EWS-FLI1 |
|
|
JQ 1 |
EWS-FLI1 |
Ewing sarcoma |
Decreased proliferation, increased apoptosis and tumor formation inhibited |
|
|
YK-4-279 |
EWS-FLI 1 |
Ewing sarcoma |
Inhibition of recruitment of co-activator molecule DHX9 for neoplastic transformation |
[276] |
|
Resveratrol |
DDX5-ETV4 |
Prostate cancer |
Degrades the DDX5 protein and inhibit mTOR pathway |
[277] |
|
JQ 1 |
TMPRSS2-ERG |
Prostate cancer |
Disrupt association between bromodomain containing domain (BRD4) with androgen receptor (AR) and inhibit transcriptional activity of AR and TMPRSS2-ERG expression |
[275] |
|
siRNA-loaded liposome |
TMPRSS2-ERG |
Prostate cancer |
Inhibition of angiogenesis and cell proliferation; Induction of apoptosis |
[278,279] |
|
siRNA |
TMPRSS2-ERG |
Prostate cancer |
Decreased cell proliferation and migration thereby metastasis; increased apoptosis |
[280] |
|
Taxol |
TMPRSS2-ERG EWS-ERG FUS-ERG |
Prostate cancer Fibroblast sarcoma |
Caused genetic instability by aneuploidy |
[281] |
|
Larotrectinib |
ETV6-NTRK3 |
Breast cancer |
Inhibition of tropomyosin kinase inhibitor (TKI) demonstrates stoppage of metastasis |
[282] |
|
Entrectinib |
ETV6-NTRK3 |
Breast cancer |
Inhibition of tropomyosin kinase inhibitor (TKI) demonstrates stoppage of metastasis |
|
|
Nutlin-3 |
ETV6-RUNX1 (E/R) |
Childhood leukemia |
Inhibits E/R induced MDM2-p53 interaction promoting cell cycle arrest and apoptosis |
[283] |
|
Valproic acid |
EWS-ERG EWS-FLI1 |
Ewing sarcoma |
Reverses the actions of fused oncoprotein on RXRα transcriptional activity and subsequently cell growth inhibited |
[284] |
|
Furamidine |
ETV6-NTRK3 |
Glioblastoma |
Binds with DNA binding domain of RBPJ (an upstream regulator ETV6-NTRK3) thereby productions of ETV6 and ETV-NTRK3 are prevented |
[122] |
Acknowledgements
Apologies to those whose related publications were not cited due to space limitations. The authors acknowledge University of Kalyani, India for supporting study. N.S. was funded by DBT- Ramalingaswami Fellowship (Award sanction No: BT/RLF/Re-entry/27/2016) from Department of Biotechnology, India. A.D. worked under a fellowship from SERB-DST (Award sanction No: ECR/2018/000595) awarded to N.S. from Department of Science and Technology, India. S.G. worked under a fellowship from UGC, India. The authors have no financial conflict of interest to report.
Authors contribution
A.D., and N.S. were responsible for conceptualization, manuscript design, data curation and graphics. S.G., A.D. and N.S. were responsible for writing and editing.
Competing Interests
The authors declare no competing interests. Correspondence and other requests regarding the work should be addressed to N.S.
References
2. Leprince D, Gegonne A, Coll J, de Taisne C, Schneeberger A, Lagrou C, et al. A putative second cell-derived oncogene of the avian leukaemia retrovirus E26. Nature. 1983;306(5941):395-7.
3. Nunn MF, Seeburg PH, Moscovici C, Duesberg PH. Tripartite structure of the avian erythroblastosis virus E26 transforming gene. Nature. 1983;306(5941):391-5.
4. Degnan BM, Degnan SM, Naganuma T, Morse DE. The ets multigene family is conserved throughout the Metazoa. Nucleic Acids Res. 1993;21(15):3479-84.
5. Karim FD, Urness LD, Thummel CS, Klemsz MJ, McKercher SR, Celada A, et al. The ETS-domain: a new DNA-binding motif that recognizes a purine-rich core DNA sequence. Genes Dev. 1990;4(9):1451-3.
6. Findlay VJ, LaRue AC, Turner DP, Watson PM, Watson DK. Understanding the role of ETS-mediated gene regulation in complex biological processes. Adv Cancer Res. 2013;119:1-61.
7. Hollenhorst PC, McIntosh LP, Graves BJ. Genomic and biochemical insights into the specificity of ETS transcription factors. Annu Rev Biochem. 2011;80:437-71.
8. Buggy Y, Maguire TM, McGreal G, McDermott E, Hill AD, O'Higgins N, et al. Overexpression of the Ets-1 transcription factor in human breast cancer. Br J Cancer. 2004;91(7):1308-15.
9. Carmichael CL, Metcalf D, Henley KJ, Kruse EA, Di Rago L, Mifsud S, et al. Hematopoietic overexpression of the transcription factor Erg induces lymphoid and erythro-megakaryocytic leukemia. Proc Natl Acad Sci U S A. 2012;109(38):15437-42.
10. Carver BS, Tran J, Gopalan A, Chen Z, Shaikh S, Carracedo A, et al. Aberrant ERG expression cooperates with loss of PTEN to promote cancer progression in the prostate. Nat Genet. 2009;41(5):619-24.
11. Fry EA, Inoue K. Aberrant expression of ETS1 and ETS2 proteins in cancer. Cancer Rep Rev. 2018;2(3).
12. Jane-Valbuena J, Widlund HR, Perner S, Johnson LA, Dibner AC, Lin WM, et al. An oncogenic role for ETV1 in melanoma. Cancer Res. 2010;70(5):2075-84.
13. Li F, Yuan Q, Di W, Xia X, Liu Z, Mao N, et al. ERG orchestrates chromatin interactions to drive prostate cell fate reprogramming. J Clin Invest. 2020;130(11):5924-41.
14. Oh S, Shin S, Song H, Grande JP, Janknecht R. Relationship between ETS Transcription Factor ETV1 and TGF-beta-regulated SMAD Proteins in Prostate Cancer. Sci Rep. 2019;9(1):8186.
15. Shimizu K, Ichikawa H, Tojo A, Kaneko Y, Maseki N, Hayashi Y, et al. An ets-related gene, ERG, is rearranged in human myeloid leukemia with t(16;21) chromosomal translocation. Proc Natl Acad Sci U S A. 1993;90(21):10280-4.
16. Smeets MF, Chan AC, Dagger S, Bradley CK, Wei A, Izon DJ. Fli-1 overexpression in hematopoietic progenitors deregulates T cell development and induces pre-T cell lymphoblastic leukaemia/lymphoma. PLoS One. 2013;8(5):e62346.
17. Truong AH, Ben-David Y. The role of Fli-1 in normal cell function and malignant transformation. Oncogene. 2000;19(55):6482-9.
18. Tsai HP, Tsai TH, Hsieh YJ, Chen YT, Lee CL, Tsai YC, et al. Overexpression of Fli-1 in astrocytoma is associated with poor prognosis. Oncotarget. 2017;8(17):29174-86.
19. Wang H, Ou Y, Ou J, Jian Z. Fli1 promotes metastasis by regulating MMP2 signaling in hepatocellular carcinoma. Mol Med Rep. 2018;17(1):1986-92.
20. Zhang L, Lemarchandel V, Romeo PH, Ben-David Y, Greer P, Bernstein A. The Fli-1 proto-oncogene, involved in erythroleukemia and Ewing's sarcoma, encodes a transcriptional activator with DNA-binding specificities distinct from other Ets family members. Oncogene. 1993;8(6):1621-30.
21. Baena E, Shao Z, Linn DE, Glass K, Hamblen MJ, Fujiwara Y, et al. ETV1 directs androgen metabolism and confers aggressive prostate cancer in targeted mice and patients. Genes Dev. 2013;27(6):683-98.
22. Becker-Santos DD, Guo Y, Ghaffari M, Vickers ED, Lehman M, Altamirano-Dimas M, et al. Integrin-linked kinase as a target for ERG-mediated invasive properties in prostate cancer models. Carcinogenesis. 2012;33(12):2558-67.
23. Birdsey GM, Dryden NH, Shah AV, Hannah R, Hall MD, Haskard DO, et al. The transcription factor Erg regulates expression of histone deacetylase 6 and multiple pathways involved in endothelial cell migration and angiogenesis. Blood. 2012;119(3):894-903.
24. Gilligan C. Adolescent development reconsidered. New Dir Child Dev. 1987(37):63-92.
25. Kruse EA, Loughran SJ, Baldwin TM, Josefsson EC, Ellis S, Watson DK, et al. Dual requirement for the ETS transcription factors Fli-1 and Erg in hematopoietic stem cells and the megakaryocyte lineage. Proc Natl Acad Sci U S A. 2009;106(33):13814-9.
26. Leshem O, Madar S, Kogan-Sakin I, Kamer I, Goldstein I, Brosh R, et al. TMPRSS2/ERG promotes epithelial to mesenchymal transition through the ZEB1/ZEB2 axis in a prostate cancer model. PLoS One. 2011;6(7):e21650.
27. Miao B, Bauer AS, Hufnagel K, Wu Y, Trajkovic-Arsic M, Pirona AC, et al. The transcription factor FLI1 promotes cancer progression by affecting cell cycle regulation. Int J Cancer. 2020;147(1):189-201.
28. Barton K, Muthusamy N, Fischer C, Ting CN, Walunas TL, Lanier LL, et al. The Ets-1 transcription factor is required for the development of natural killer cells in mice. Immunity. 1998;9(4):555-63.
29. Muthusamy N, Barton K, Leiden JM. Defective activation and survival of T cells lacking the Ets-1 transcription factor. Nature. 1995;377(6550):639-42.
30. Ramirez K, Chandler KJ, Spaulding C, Zandi S, Sigvardsson M, Graves BJ, et al. Gene deregulation and chronic activation in natural killer cells deficient in the transcription factor ETS1. Immunity. 2012;36(6):921-32.
31. Sunshine A, Goich D, Stith A, Sortino K, Dalton J, Metcalfe S, et al. Ets1 Controls the Development of B Cell Autoimmune Responses in a Cell-Intrinsic Manner. Immunohorizons. 2019;3(7):331-40.
32. Wei G, Srinivasan R, Cantemir-Stone CZ, Sharma SM, Santhanam R, Weinstein M, et al. Ets1 and Ets2 are required for endothelial cell survival during embryonic angiogenesis. Blood. 2009;114(5):1123-30.
33. Ye M, Xu L, Fu M, Chen D, Mattina T, Zufardi O, et al. Gene-targeted deletion in mice of the Ets-1 transcription factor, a candidate gene in the Jacobsen syndrome kidney "critical region," causes abnormal kidney development. Am J Med Genet A. 2019;179(1):71-7.
34. Zhan Y, Brown C, Maynard E, Anshelevich A, Ni W, Ho IC, et al. Ets-1 is a critical regulator of Ang II-mediated vascular inflammation and remodeling. J Clin Invest. 2005;115(9):2508-16.
35. Bartel FO, Higuchi T, Spyropoulos DD. Mouse models in the study of the Ets family of transcription factors. Oncogene. 2000;19(55):6443-54.
36. Yamamoto H, Flannery ML, Kupriyanov S, Pearce J, McKercher SR, Henkel GW, et al. Defective trophoblast function in mice with a targeted mutation of Ets2. Genes Dev. 1998;12(9):1315-26.
37. Zaldumbide A, Carlotti F, Pognonec P, Boulukos KE. The role of the Ets2 transcription factor in the proliferation, maturation, and survival of mouse thymocytes. J Immunol. 2002;169(9):4873-81.
38. Birdsey GM, Shah AV, Dufton N, Reynolds LE, Osuna Almagro L, Yang Y, et al. The endothelial transcription factor ERG promotes vascular stability and growth through Wnt/beta-catenin signaling. Dev Cell. 2015;32(1):82-96.
39. Rommel C, Rosner S, Lother A, Barg M, Schwaderer M, Gilsbach R, et al. The Transcription Factor ETV1 Induces Atrial Remodeling and Arrhythmia. Circ Res. 2018;123(5):550-63.
40. Das S, Koyano-Nakagawa N, Gafni O, Maeng G, Singh BN, Rasmussen T, et al. Generation of human endothelium in pig embryos deficient in ETV2. Nat Biotechnol. 2020;38(3):297-302.
41. Garry DJ. Etv2 IS A MASTER REGULATOR OF HEMATOENDOTHELIAL LINEAGES. Trans Am Clin Climatol Assoc. 2016;127:212-23.
42. Koyano-Nakagawa N, Garry DJ. Etv2 as an essential regulator of mesodermal lineage development. Cardiovasc Res. 2017;113(11):1294-306.
43. Liu F, Li D, Yu YY, Kang I, Cha MJ, Kim JY, et al. Induction of hematopoietic and endothelial cell program orchestrated by ETS transcription factor ER71/ETV2. EMBO Rep. 2015;16(5):654-69.
44. Singh BN, Gong W, Das S, Theisen JWM, Sierra-Pagan JE, Yannopoulos D, et al. Etv2 transcriptionally regulates Yes1 and promotes cell proliferation during embryogenesis. Sci Rep. 2019;9(1):9736.
45. Rodriguez AC, Vahrenkamp JM, Berrett KC, Clark KA, Guillen KP, Scherer SD, et al. ETV4 Is Necessary for Estrogen Signaling and Growth in Endometrial Cancer Cells. Cancer Res. 2020;80(6):1234-45.
46. Zhang X, Zhao X, Li G, Zhang M, Xing P, Li Z, et al. Establishment of Etv5 gene knockout mice as a recipient model for spermatogonial stem cell transplantation. Biol Open. 2021;10(1).
47. Zhang Z, Newton K, Kummerfeld SK, Webster J, Kirkpatrick DS, Phu L, et al. Transcription factor Etv5 is essential for the maintenance of alveolar type II cells. Proc Natl Acad Sci U S A. 2017;114(15):3903-8.
48. Cesari F, Rennekampff V, Vintersten K, Vuong LG, Seibler J, Bode J, et al. Elk-1 knock-out mice engineered by Flp recombinase-mediated cassette exchange. Genesis. 2004;38(2):87-92.
49. Maurice D, Costello P, Sargent M, Treisman R. ERK Signaling Controls Innate-like CD8(+) T Cell Differentiation via the ELK4 (SAP-1) and ELK1 Transcription Factors. J Immunol. 2018;201(6):1681-91.
50. Chen YH, Layne MD, Chung SW, Ejima K, Baron RM, Yet SF, et al. Elk-3 is a transcriptional repressor of nitric-oxide synthase 2. J Biol Chem. 2003;278(41):39572-7.
51. Weinl C, Wasylyk C, Garcia Garrido M, Sothilingam V, Beck SC, Riehle H, et al. Elk3 deficiency causes transient impairment in post-natal retinal vascular development and formation of tortuous arteries in adult murine retinae. PLoS One. 2014;9(9):e107048.
52. Paulsson JO, Wang N, Gao J, Stenman A, Zedenius J, Mu N, et al. GABPA-dependent down-regulation of DICER1 in follicular thyroid tumours. Endocr Relat Cancer. 2020;27(5):295-308.
53. Yuan X, Mu N, Wang N, Straat K, Sofiadis A, Guo Y, et al. GABPA inhibits invasion/metastasis in papillary thyroid carcinoma by regulating DICER1 expression. Oncogene. 2019;38(7):965-79.
54. Larsen S, Kawamoto S, Tanuma S, Uchiumi F. The hematopoietic regulator, ELF-1, enhances the transcriptional response to Interferon-beta of the OAS1 anti-viral gene. Sci Rep. 2015;5:17497.
55. Sun G, Qiu L, Yu L, An Y, Ding Y, Zhou L, et al. Loss of Function Mutation in ELF4 Causes Autoinflammatory and Immunodeficiency Disease in Human. J Clin Immunol. 2022;42(4):798-810.
56. Rasighaemi P, Onnebo SM, Liongue C, Ward AC. ETV6 (TEL1) regulates embryonic hematopoiesis in zebrafish. Haematologica. 2015;100(1):23-31.
57. Schotte R, Nagasawa M, Weijer K, Spits H, Blom B. The ETS transcription factor Spi-B is required for human plasmacytoid dendritic cell development. J Exp Med. 2004;200(11):1503-9.
58. Alam Z, Devalaraja S, Li M, To TKJ, Folkert IW, Mitchell-Velasquez E, et al. Counter Regulation of Spic by NF-kappaB and STAT Signaling Controls Inflammation and Iron Metabolism in Macrophages. Cell Rep. 2020;31(13):107825.
59. Moreau-Gachelin F, Wendling F, Molina T, Denis N, Titeux M, Grimber G, et al. Spi-1/PU.1 transgenic mice develop multistep erythroleukemias. Mol Cell Biol. 1996;16(5):2453-63.
60. Si L, Yang R, Liu J, Dong Y, Zhang H, Xu X. Generation of two ERF gene knockout human embryonic stem cell lines using CRISPR/Cas9 system. Stem Cell Res. 2019;41:101644.
61. Ng AY, Waring P, Ristevski S, Wang C, Wilson T, Pritchard M, et al. Inactivation of the transcription factor Elf3 in mice results in dysmorphogenesis and altered differentiation of intestinal epithelium. Gastroenterology. 2002;122(5):1455-66.
62. Choi YS, Chakrabarti R, Escamilla-Hernandez R, Sinha S. Elf5 conditional knockout mice reveal its role as a master regulator in mammary alveolar development: failure of Stat5 activation and functional differentiation in the absence of Elf5. Dev Biol. 2009;329(2):227-41.
63. Marko CK, Menon BB, Chen G, Whitsett JA, Clevers H, Gipson IK. Spdef null mice lack conjunctival goblet cells and provide a model of dry eye. Am J Pathol. 2013;183(1):35-48.
64. Bradner JE, Hnisz D, Young RA. Transcriptional Addiction in Cancer. Cell. 2017;168(4):629-43.
65. Herz HM. Enhancer deregulation in cancer and other diseases. Bioessays. 2016;38(10):1003-15.
66. Huang T, Li J, Wang SM. Core promoter mutation contributes to abnormal gene expression in bladder cancer. BMC Cancer. 2022;22(1):68.
67. Sharma S, Kelly TK, Jones PA. Epigenetics in cancer. Carcinogenesis. 2010;31(1):27-36.
68. Majerus MA, Bibollet-Ruche F, Telliez JB, Wasylyk B, Bailleul B. Serum, AP-1 and Ets-1 stimulate the human ets-1 promoter. Nucleic Acids Res. 1992;20(11):2699-703.
69. Oikawa M, Abe M, Kurosawa H, Hida W, Shirato K, Sato Y. Hypoxia induces transcription factor ETS-1 via the activity of hypoxia-inducible factor-1. Biochem Biophys Res Commun. 2001;289(1):39-43.
70. Seth A, Papas TS. The c-ets-1 proto-oncogene has oncogenic activity and is positively autoregulated. Oncogene. 1990;5(12):1761-7.
71. Goetze S, Kintscher U, Kaneshiro K, Meehan WP, Collins A, Fleck E, et al. TNFalpha induces expression of transcription factors c-fos, Egr-1, and Ets-1 in vascular lesions through extracellular signal-regulated kinases 1/2. Atherosclerosis. 2001;159(1):93-101.
72. Paumelle R, Tulasne D, Kherrouche Z, Plaza S, Leroy C, Reveneau S, et al. Hepatocyte growth factor/scatter factor activates the ETS1 transcription factor by a RAS-RAF-MEK-ERK signaling pathway. Oncogene. 2002;21(15):2309-19.
73. Igarashi T, Abe M, Oikawa M, Nukiwa T, Sato Y. Retinoic acids repress the expression of ETS-1 in endothelial cells. Tohoku J Exp Med. 2001;194(1):35-43.
74. Raouf A, Li V, Kola I, Watson DK, Seth A. The Ets1 proto-oncogene is upregulated by retinoic acid: characterization of a functional retinoic acid response element in the Ets1 promoter. Oncogene. 2000;19(15):1969-74.
75. So EN, Crowe DL. Characterization of a retinoic acid responsive element in the human ets-1 promoter. IUBMB Life. 2000;50(6):365-70.
76. Kim GC, Kwon HK, Lee CG, Verma R, Rudra D, Kim T, et al. Upregulation of Ets1 expression by NFATc2 and NFKB1/RELA promotes breast cancer cell invasiveness. Oncogenesis. 2018;7(11):91.
77. Iotsova V, Crepieux P, Montpellier C, Laudet V, Stehelin D. TATA-less promoters of some Ets-family genes are efficiently repressed by wild-type p53. Oncogene. 1996;13(11):2331-7.
78. Sgouras DN, Athanasiou MA, Beal GJ, Jr., Fisher RJ, Blair DG, Mavrothalassitis GJ. ERF: an ETS domain protein with strong transcriptional repressor activity, can suppress ets-associated tumorigenesis and is regulated by phosphorylation during cell cycle and mitogenic stimulation. EMBO J. 1995;14(19):4781-93.
79. Wang DM, Sevcikova S, Wen H, Roberts S, Lipsick JS. v-Myb represses the transcription of Ets-2. Oncogene. 2007;26(8):1238-44.
80. Adamo P, Ladomery MR. The oncogene ERG: a key factor in prostate cancer. Oncogene. 2016;35(4):403-14.
81. Bohne A, Schlee C, Mossner M, Thibaut J, Heesch S, Thiel E, et al. Epigenetic control of differential expression of specific ERG isoforms in acute T-lymphoblastic leukemia. Leuk Res. 2009;33(6):817-22.
82. Schwartzman J, Mongoue-Tchokote S, Gibbs A, Gao L, Corless CL, Jin J, et al. A DNA methylation microarray-based study identifies ERG as a gene commonly methylated in prostate cancer. Epigenetics. 2011;6(10):1248-56.
83. Ginsberg JP, de Alava E, Ladanyi M, Wexler LH, Kovar H, Paulussen M, et al. EWS-FLI1 and EWS-ERG gene fusions are associated with similar clinical phenotypes in Ewing's sarcoma. J Clin Oncol. 1999;17(6):1809-14.
84. Giovannini M, Biegel JA, Serra M, Wang JY, Wei YH, Nycum L, et al. EWS-erg and EWS-Fli1 fusion transcripts in Ewing's sarcoma and primitive neuroectodermal tumors with variant translocations. J Clin Invest. 1994;94(2):489-96.
85. Ichikawa H, Shimizu K, Hayashi Y, Ohki M. An RNA-binding protein gene, TLS/FUS, is fused to ERG in human myeloid leukemia with t(16;21) chromosomal translocation. Cancer Res. 1994;54(11):2865-8.
86. Pflueger D, Rickman DS, Sboner A, Perner S, LaFargue CJ, Svensson MA, et al. N-myc downstream regulated gene 1 (NDRG1) is fused to ERG in prostate cancer. Neoplasia. 2009;11(8):804-11.
87. Sorensen PH, Lessnick SL, Lopez-Terrada D, Liu XF, Triche TJ, Denny CT. A second Ewing's sarcoma translocation, t(21;22), fuses the EWS gene to another ETS-family transcription factor, ERG. Nat Genet. 1994;6(2):146-51.
88. Sotoca AM, Prange KH, Reijnders B, Mandoli A, Nguyen LN, Stunnenberg HG, et al. The oncofusion protein FUS-ERG targets key hematopoietic regulators and modulates the all-trans retinoic acid signaling pathway in t(16;21) acute myeloid leukemia. Oncogene. 2016;35(15):1965-76.
89. Tomlins SA, Laxman B, Varambally S, Cao X, Yu J, Helgeson BE, et al. Role of the TMPRSS2-ERG gene fusion in prostate cancer. Neoplasia. 2008;10(2):177-88.
90. Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun XW, et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science. 2005;310(5748):644-8.
91. Mani RS, Iyer MK, Cao Q, Brenner JC, Wang L, Ghosh A, et al. TMPRSS2-ERG-mediated feed-forward regulation of wild-type ERG in human prostate cancers. Cancer Res. 2011;71(16):5387-92.
92. May WA, Lessnick SL, Braun BS, Klemsz M, Lewis BC, Lunsford LB, et al. The Ewing's sarcoma EWS/FLI-1 fusion gene encodes a more potent transcriptional activator and is a more powerful transforming gene than FLI-1. Mol Cell Biol. 1993;13(12):7393-8.
93. Kwiatkowski BA, Zielinska-Kwiatkowska AG, Bauer TR, Jr., Hickstein DD. The ETS family member Tel antagonizes the Fli-1 phenotype in hematopoietic cells. Blood Cells Mol Dis. 2000;26(1):84-90.
94. Lelievre E, Lionneton F, Mattot V, Spruyt N, Soncin F. Ets-1 regulates fli-1 expression in endothelial cells. Identification of ETS binding sites in the fli-1 gene promoter. J Biol Chem. 2002;277(28):25143-51.
95. Svenson JL, Chike-Harris K, Amria MY, Nowling TK. The mouse and human Fli1 genes are similarly regulated by Ets factors in T cells. Genes Immun. 2010;11(2):161-72.
96. Abedin MJ, Nguyen A, Jiang N, Perry CE, Shelton JM, Watson DK, et al. Fli1 acts downstream of Etv2 to govern cell survival and vascular homeostasis via positive autoregulation. Circ Res. 2014;114(11):1690-9.
97. Krueger KC, Deneris ES. Serotonergic transcription of human FEV reveals direct GATA factor interactions and fate of Pet-1-deficient serotonin neuron precursors. J Neurosci. 2008;28(48):12748-58.
98. Liang YX, Liang YK, Zou ZH, Zhuo YJ, Ye JH, Zhu XJ, et al. Tumor Suppressor Role and Clinical Significance of the FEV Gene in Prostate Cancer. Dis Markers. 2022;2022:8724035.
99. Boone MA, Taslim C, Crow JC, Selich-Anderson J, Watson M, Heppner P, et al. Identification of a Novel FUS/ETV4 Fusion and Comparative Analysis with Other Ewing Sarcoma Fusion Proteins. Mol Cancer Res. 2021;19(11):1795-801.
100. Ng TL, O'Sullivan MJ, Pallen CJ, Hayes M, Clarkson PW, Winstanley M, et al. Ewing sarcoma with novel translocation t(2;16) producing an in-frame fusion of FUS and FEV. J Mol Diagn. 2007;9(4):459-63.
101. Li Q, Meissner TB, Wang F, Du Z, Ma S, Kshirsagar S, et al. ELF3 activated by a superenhancer and an autoregulatory feedback loop is required for high-level HLA-C expression on extravillous trophoblasts. Proc Natl Acad Sci U S A. 2021;118(9).
102. Enfield KSS, Marshall EA, Anderson C, Ng KW, Rahmati S, Xu Z, et al. Epithelial tumor suppressor ELF3 is a lineage-specific amplified oncogene in lung adenocarcinoma. Nat Commun. 2019;10(1):5438.
103. Chang CH, Scott GK, Kuo WL, Xiong X, Suzdaltseva Y, Park JW, et al. ESX: a structurally unique Ets overexpressed early during human breast tumorigenesis. Oncogene. 1997;14(13):1617-22.
104. Eckel KL, Tentler JJ, Cappetta GJ, Diamond SE, Gutierrez-Hartmann A. The epithelial-specific ETS transcription factor ESX/ESE-1/Elf-3 modulates breast cancer-associated gene expression. DNA Cell Biol. 2003;22(2):79-94.
105. Han W, Zhang C, Shi CT, Gao XJ, Zhou MH, Shao QX, et al. Roles of eIF3m in the tumorigenesis of triple negative breast cancer. Cancer Cell Int. 2020;20:141.
106. Yeung TL, Leung CS, Wong KK, Gutierrez-Hartmann A, Kwong J, Gershenson DM, et al. ELF3 is a negative regulator of epithelial-mesenchymal transition in ovarian cancer cells. Oncotarget. 2017;8(10):16951-63.
107. Zheng L, Xu M, Xu J, Wu K, Fang Q, Liang Y, et al. ELF3 promotes epithelial-mesenchymal transition by protecting ZEB1 from miR-141-3p-mediated silencing in hepatocellular carcinoma. Cell Death Dis. 2018;9(3):387.
108. Lee HJ, Hinshelwood RA, Bouras T, Gallego-Ortega D, Valdes-Mora F, Blazek K, et al. Lineage specific methylation of the Elf5 promoter in mammary epithelial cells. Stem Cells. 2011;29(10):1611-9.
109. Ng RK, Dean W, Dawson C, Lucifero D, Madeja Z, Reik W, et al. Epigenetic restriction of embryonic cell lineage fate by methylation of Elf5. Nat Cell Biol. 2008;10(11):1280-90.
110. Zhou J, Chehab R, Tkalcevic J, Naylor MJ, Harris J, Wilson TJ, et al. Elf5 is essential for early embryogenesis and mammary gland development during pregnancy and lactation. EMBO J. 2005;24(3):635-44.
111. Piggin CL, Roden DL, Gallego-Ortega D, Lee HJ, Oakes SR, Ormandy CJ. ELF5 isoform expression is tissue-specific and significantly altered in cancer. Breast Cancer Res. 2016;18(1):4.
112. Dittmer J. Nuclear Mechanisms Involved in Endocrine Resistance. Front Oncol. 2021;11:736597.
113. Kalyuga M, Gallego-Ortega D, Lee HJ, Roden DL, Cowley MJ, Caldon CE, et al. ELF5 suppresses estrogen sensitivity and underpins the acquisition of antiestrogen resistance in luminal breast cancer. PLoS Biol. 2012;10(12):e1001461.
114. Chakrabarti R, Hwang J, Andres Blanco M, Wei Y, Lukacisin M, Romano RA, et al. Elf5 inhibits the epithelial-mesenchymal transition in mammary gland development and breast cancer metastasis by transcriptionally repressing Snail2. Nat Cell Biol. 2012;14(11):1212-22.
115. Qu X, Li Q, Tu S, Yang X, Wen W. ELF5 inhibits the proliferation and invasion of breast cancer cells by regulating CD24. Mol Biol Rep. 2021;48(6):5023-32.
116. Barjesteh van Waalwijk van Doorn-Khosrovani S, Spensberger D, de Knegt Y, Tang M, Lowenberg B, Delwel R. Somatic heterozygous mutations in ETV6 (TEL) and frequent absence of ETV6 protein in acute myeloid leukemia. Oncogene. 2005;24(25):4129-37.
117. Tognon C, Knezevich SR, Huntsman D, Roskelley CD, Melnyk N, Mathers JA, et al. Expression of the ETV6-NTRK3 gene fusion as a primary event in human secretory breast carcinoma. Cancer Cell. 2002;2(5):367-76.
118. Wang M, Gu D, Du M, Xu Z, Zhang S, Zhu L, et al. Common genetic variation in ETV6 is associated with colorectal cancer susceptibility. Nat Commun. 2016;7:11478.
119. Di Paola J, Porter CC. ETV6-related thrombocytopenia and leukemia predisposition. Blood. 2019;134(8):663-7.
120. Keller M, Hopp L, Liu X, Wohland T, Rohde K, Cancello R, et al. Genome-wide DNA promoter methylation and transcriptome analysis in human adipose tissue unravels novel candidate genes for obesity. Mol Metab. 2017;6(1):86-100.
121. Neveu B, Richer C, Cassart P, Caron M, Jimenez-Cortes C, St-Onge P, et al. Identification of new ETV6 modulators through a high-throughput functional screening. iScience. 2022;25(3):103858.
122. Biswas A, Rajesh Y, Das S, Banerjee I, Kapoor N, Mitra P, et al. Therapeutic targeting of RBPJ, an upstream regulator of ETV6 gene, abrogates ETV6-NTRK3 fusion gene transformations in glioblastoma. Cancer Lett. 2022;544:215811.
123. Biswas A, Rajesh Y, Mitra P, Mandal M. ETV6 gene aberrations in non-haematological malignancies: A review highlighting ETV6 associated fusion genes in solid tumors. Biochim Biophys Acta Rev Cancer. 2020;1874(1):188389.
124. De Braekeleer E, Douet-Guilbert N, Morel F, Le Bris MJ, Basinko A, De Braekeleer M. ETV6 fusion genes in hematological malignancies: a review. Leuk Res. 2012;36(8):945-61.
125. Leeman-Neill RJ, Kelly LM, Liu P, Brenner AV, Little MP, Bogdanova TI, et al. ETV6-NTRK3 is a common chromosomal rearrangement in radiation-associated thyroid cancer. Cancer. 2014;120(6):799-807.
126. Peeters P, Raynaud SD, Cools J, Wlodarska I, Grosgeorge J, Philip P, et al. Fusion of TEL, the ETS-variant gene 6 (ETV6), to the receptor-associated kinase JAK2 as a result of t(9;12) in a lymphoid and t(9;15;12) in a myeloid leukemia. Blood. 1997;90(7):2535-40.
127. Schwaller J. Modeling ETV6-JAK2-induced leukemia: insights from the zebrafish. Haematologica. 2012;97(12):1783-5.
128. Xu LS, Francis A, Turkistany S, Shukla D, Wong A, Batista CR, et al. ETV6-RUNX1 interacts with a region in SPIB intron 1 to regulate gene expression in pre-B-cell acute lymphoblastic leukemia. Exp Hematol. 2019;73:50-63 e2.
129. Calero-Nieto FJ, Wood AD, Wilson NK, Kinston S, Landry JR, Gottgens B. Transcriptional regulation of Elf-1: locus-wide analysis reveals four distinct promoters, a tissue-specific enhancer, control by PU.1 and the importance of Elf-1 downregulation for erythroid maturation. Nucleic Acids Res. 2010;38(19):6363-74.
130. Wang CY, Petryniak B, Thompson CB, Kaelin WG, Leiden JM. Regulation of the Ets-related transcription factor Elf-1 by binding to the retinoblastoma protein. Science. 1993;260(5112):1330-5.
131. Budka JA, Ferris MW, Capone MJ, Hollenhorst PC. Common ELF1 deletion in prostate cancer bolsters oncogenic ETS function, inhibits senescence and promotes docetaxel resistance. Genes Cancer. 2018;9(5-6):198-214.
132. Cheng M, Zeng Y, Zhang T, Xu M, Li Z, Wu Y. Transcription Factor ELF1 Activates MEIS1 Transcription and Then Regulates the GFI1/FBW7 Axis to Promote the Development of Glioma. Mol Ther Nucleic Acids. 2021;23:418-30.
133. Qiu Y, Morii E, Zhang B, Tomita Y, Aozasa K. E74-like factor 2 transactivates valosin-containing protein gene, a gene involved in cancer growth. Exp Mol Pathol. 2008;84(3):226-9.
134. Wilkinson DA, Neale GA, Mao S, Naeve CW, Goorha RM. Elf-2, a rhombotin-2 binding ets transcription factor: discovery and potential role in T cell leukemia. Leukemia. 1997;11(1):86-96.
135. Oh S, Shin S, Janknecht R. ETV1, 4 and 5: an oncogenic subfamily of ETS transcription factors. Biochim Biophys Acta. 2012;1826(1):1-12.
136. Abe H, Okazawa M, Nakanishi S. The Etv1/Er81 transcription factor orchestrates activity-dependent gene regulation in the terminal maturation program of cerebellar granule cells. Proc Natl Acad Sci U S A. 2011;108(30):12497-502.
137. Cai C, Hsieh CL, Omwancha J, Zheng Z, Chen SY, Baert JL, et al. ETV1 is a novel androgen receptor-regulated gene that mediates prostate cancer cell invasion. Mol Endocrinol. 2007;21(8):1835-46.
138. Oh S, Song H, Freeman WM, Shin S, Janknecht R. Cooperation between ETS transcription factor ETV1 and histone demethylase JMJD1A in colorectal cancer. Int J Oncol. 2020;57(6):1319-32.
139. Shin S, Bosc DG, Ingle JN, Spelsberg TC, Janknecht R. Rcl is a novel ETV1/ER81 target gene upregulated in breast tumors. J Cell Biochem. 2008;105(3):866-74.
140. Shin S, Kim TD, Jin F, van Deursen JM, Dehm SM, Tindall DJ, et al. Induction of prostatic intraepithelial neoplasia and modulation of androgen receptor by ETS variant 1/ETS-related protein 81. Cancer Res. 2009;69(20):8102-10.
141. Barros-Silva JD, Paulo P, Bakken AC, Cerveira N, Lovf M, Henrique R, et al. Novel 5' fusion partners of ETV1 and ETV4 in prostate cancer. Neoplasia. 2013;15(7):720-6.
142. Raskin GA, Pozharisski KM, Iyevleva AG, Rikov IV, Orlova RV, Imyanitov EN. Unusual Clinical Presentation of Gastrointestinal Clear Cell Sarcoma. Gastrointest Tumors. 2015;2(2):83-8.
143. Dumortier M, Ladam F, Damour I, Vacher S, Bieche I, Marchand N, et al. ETV4 transcription factor and MMP13 metalloprotease are interplaying actors of breast tumorigenesis. Breast Cancer Res. 2018;20(1):73.
144. Tyagi N, Deshmukh SK, Srivastava SK, Azim S, Ahmad A, Al-Ghadhban A, et al. ETV4 Facilitates Cell-Cycle Progression in Pancreatic Cells through Transcriptional Regulation of Cyclin D1. Mol Cancer Res. 2018;16(2):187-96.
145. Yang L, Fox SA, Kirby JL, Troan BV, Hinton BT. Putative regulation of expression of members of the Ets variant 4 transcription factor family and their downstream targets in the rat epididymis. Biol Reprod. 2006;74(4):714-20.
146. Zheng C, Liu M, Ge Y, Qian Y, Fan H. HBx increases chromatin accessibility and ETV4 expression to regulate dishevelled-2 and promote HCC progression. Cell Death Dis. 2022;13(2):116.
147. Han B, Mehra R, Dhanasekaran SM, Yu J, Menon A, Lonigro RJ, et al. A fluorescence in situ hybridization screen for E26 transformation-specific aberrations: identification of DDX5-ETV4 fusion protein in prostate cancer. Cancer Res. 2008;68(18):7629-37.
148. Panagopoulos I, Gorunova L, Viset T, Heim S. Gene fusions AHRR-NCOA2, NCOA2-ETV4, ETV4-AHRR, P4HA2-TBCK, and TBCK-P4HA2 resulting from the translocations t(5;8;17)(p15;q13;q21) and t(4;5)(q24;q31) in a soft tissue angiofibroma. Oncol Rep. 2016;36(5):2455-62.
149. Tomlins SA, Mehra R, Rhodes DR, Smith LR, Roulston D, Helgeson BE, et al. TMPRSS2:ETV4 gene fusions define a third molecular subtype of prostate cancer. Cancer Res. 2006;66(7):3396-400.
150. Mus LM, Lambertz I, Claeys S, Kumps C, Van Loocke W, Van Neste C, et al. The ETS transcription factor ETV5 is a target of activated ALK in neuroblastoma contributing to increased tumour aggressiveness. Sci Rep. 2020;10(1):218.
151. Giri AK. Higher ETV5 Expression Associates With Poor 5-Florouracil-Based Adjuvant Therapy Response in Colon Cancer. Front Pharmacol. 2020;11:620811.
152. di Martino E, Alder O, Hurst CD, Knowles MA. ETV5 links the FGFR3 and Hippo signalling pathways in bladder cancer. Sci Rep. 2019;9(1):5740.
153. Llaurado M, Majem B, Castellvi J, Cabrera S, Gil-Moreno A, Reventos J, et al. Analysis of gene expression regulated by the ETV5 transcription factor in OV90 ovarian cancer cells identifies FOXM1 overexpression in ovarian cancer. Mol Cancer Res. 2012;10(7):914-24.
154. Monge M, Colas E, Doll A, Gonzalez M, Gil-Moreno A, Planaguma J, et al. ERM/ETV5 up-regulation plays a role during myometrial infiltration through matrix metalloproteinase-2 activation in endometrial cancer. Cancer Res. 2007;67(14):6753-9.
155. Helgeson BE, Tomlins SA, Shah N, Laxman B, Cao Q, Prensner JR, et al. Characterization of TMPRSS2:ETV5 and SLC45A3:ETV5 gene fusions in prostate cancer. Cancer Res. 2008;68(1):73-80.
156. Kawahara T, Shareef HK, Aljarah AK, Ide H, Li Y, Kashiwagi E, et al. ELK1 is up-regulated by androgen in bladder cancer cells and promotes tumor progression. Oncotarget. 2015;6(30):29860-76.
157. Sgambato V, Vanhoutte P, Pages C, Rogard M, Hipskind R, Besson MJ, et al. In vivo expression and regulation of Elk-1, a target of the extracellular-regulated kinase signaling pathway, in the adult rat brain. J Neurosci. 1998;18(1):214-26.
158. Buchwalter G, Gross C, Wasylyk B. Ets ternary complex transcription factors. Gene. 2004;324:1-14.
159. Semenchenko K, Wasylyk C, Cheung H, Tourrette Y, Maas P, Schalken JA, et al. XRP44X, an Inhibitor of Ras/Erk Activation of the Transcription Factor Elk3, Inhibits Tumour Growth and Metastasis in Mice. PLoS One. 2016;11(7):e0159531.
160. Yoo SM, Lee CJ, An HJ, Lee JY, Lee HS, Kang HC, et al. RSK2-Mediated ELK3 Activation Enhances Cell Transformation and Breast Cancer Cell Growth by Regulation of c-fos Promoter Activity. Int J Mol Sci. 2019;20(8).
161. Gross C, Buchwalter G, Dubois-Pot H, Cler E, Zheng H, Wasylyk B. The ternary complex factor net is downregulated by hypoxia and regulates hypoxia-responsive genes. Mol Cell Biol. 2007;27(11):4133-41.
162. Robertson ED, Wasylyk C, Ye T, Jung AC, Wasylyk B. The oncogenic MicroRNA Hsa-miR-155-5p targets the transcription factor ELK3 and links it to the hypoxia response. PLoS One. 2014;9(11):e113050.
163. Rickman DS, Pflueger D, Moss B, VanDoren VE, Chen CX, de la Taille A, et al. SLC45A3-ELK4 is a novel and frequent erythroblast transformation-specific fusion transcript in prostate cancer. Cancer Res. 2009;69(7):2734-8.
164. Day BW, Stringer BW, Spanevello MD, Charmsaz S, Jamieson PR, Ensbey KS, et al. ELK4 neutralization sensitizes glioblastoma to apoptosis through downregulation of the anti-apoptotic protein Mcl-1. Neuro Oncol. 2011;13(11):1202-12.
165. Zheng L, Xu H, Di Y, Chen L, Liu J, Kang L, et al. ELK4 promotes the development of gastric cancer by inducing M2 polarization of macrophages through regulation of the KDM5A-PJA2-KSR1 axis. J Transl Med. 2021;19(1):342.
166. Baker DA, Mille-Baker B, Wainwright SM, Ish-Horowicz D, Dibb NJ. Mae mediates MAP kinase phosphorylation of Ets transcription factors in Drosophila. Nature. 2001;411(6835):330-4.
167. Kim CA, Phillips ML, Kim W, Gingery M, Tran HH, Robinson MA, et al. Polymerization of the SAM domain of TEL in leukemogenesis and transcriptional repression. EMBO J. 2001;20(15):4173-82.
168. Lacronique V, Boureux A, Valle VD, Poirel H, Quang CT, Mauchauffe M, et al. A TEL-JAK2 fusion protein with constitutive kinase activity in human leukemia. Science. 1997;278(5341):1309-12.
169. Hassler M, Richmond TJ. The B-box dominates SAP-1-SRF interactions in the structure of the ternary complex. EMBO J. 2001;20(12):3018-28.
170. Jeffery HM, Weinzierl ROJ. Multivalent and Bidirectional Binding of Transcriptional Transactivation Domains to the MED25 Coactivator. Biomolecules. 2020;10(9).
171. Shore P, Sharrocks AD. The transcription factors Elk-1 and serum response factor interact by direct protein-protein contacts mediated by a short region of Elk-1. Mol Cell Biol. 1994;14(5):3283-91.
172. Yang SH, Bumpass DC, Perkins ND, Sharrocks AD. The ETS domain transcription factor Elk-1 contains a novel class of repression domain. Mol Cell Biol. 2002;22(14):5036-46.
173. Yang SH, Shore P, Willingham N, Lakey JH, Sharrocks AD. The mechanism of phosphorylation-inducible activation of the ETS-domain transcription factor Elk-1. EMBO J. 1999;18(20):5666-74.
174. Charlot C, Dubois-Pot H, Serchov T, Tourrette Y, Wasylyk B. A review of post-translational modifications and subcellular localization of Ets transcription factors: possible connection with cancer and involvement in the hypoxic response. Methods Mol Biol. 2010;647:3-30.
175. Kedage V, Strittmatter BG, Dausinas PB, Hollenhorst PC. Phosphorylation of the oncogenic transcription factor ERG in prostate cells dissociates polycomb repressive complex 2, allowing target gene activation. J Biol Chem. 2017;292(42):17225-35.
176. Selvaraj N, Budka JA, Ferris MW, Jerde TJ, Hollenhorst PC. Prostate cancer ETS rearrangements switch a cell migration gene expression program from RAS/ERK to PI3K/AKT regulation. Mol Cancer. 2014;13:61.
177. Strittmatter BG, Jerde TJ, Hollenhorst PC. Ras/ERK and PI3K/AKT signaling differentially regulate oncogenic ERG mediated transcription in prostate cells. PLoS Genet. 2021;17(7):e1009708.
178. Huang Y, Thoms JA, Tursky ML, Knezevic K, Beck D, Chandrakanthan V, et al. MAPK/ERK2 phosphorylates ERG at serine 283 in leukemic cells and promotes stem cell signatures and cell proliferation. Leukemia. 2016;30(7):1552-61.
179. Selvaraj N, Kedage V, Hollenhorst PC. Comparison of MAPK specificity across the ETS transcription factor family identifies a high-affinity ERK interaction required for ERG function in prostate cells. Cell Commun Signal. 2015;13:12.
180. Foulds CE, Nelson ML, Blaszczak AG, Graves BJ. Ras/mitogen-activated protein kinase signaling activates Ets-1 and Ets-2 by CBP/p300 recruitment. Mol Cell Biol. 2004;24(24):10954-64.
181. Hollenhorst PC. RAS/ERK pathway transcriptional regulation through ETS/AP-1 binding sites. Small GTPases. 2012;3(3):154-8.
182. Plotnik JP, Budka JA, Ferris MW, Hollenhorst PC. ETS1 is a genome-wide effector of RAS/ERK signaling in epithelial cells. Nucleic Acids Res. 2014;42(19):11928-40.
183. Dittmer J. The role of the transcription factor Ets1 in carcinoma. Semin Cancer Biol. 2015;35:20-38.
184. Nelson ML, Kang HS, Lee GM, Blaszczak AG, Lau DK, McIntosh LP, et al. Ras signaling requires dynamic properties of Ets1 for phosphorylation-enhanced binding to coactivator CBP. Proc Natl Acad Sci U S A. 2010;107(22):10026-31.
185. Sinh ND, Endo K, Miyazawa K, Saitoh M. Ets1 and ESE1 reciprocally regulate expression of ZEB1/ZEB2, dependent on ERK1/2 activity, in breast cancer cells. Cancer Sci. 2017;108(5):952-60.
186. Yordy JS, Muise-Helmericks RC. Signal transduction and the Ets family of transcription factors. Oncogene. 2000;19(55):6503-13.
187. Svensson S, Jirstrom K, Ryden L, Roos G, Emdin S, Ostrowski MC, et al. ERK phosphorylation is linked to VEGFR2 expression and Ets-2 phosphorylation in breast cancer and is associated with tamoxifen treatment resistance and small tumours with good prognosis. Oncogene. 2005;24(27):4370-9.
188. Booy EP, Henson ES, Gibson SB. Epidermal growth factor regulates Mcl-1 expression through the MAPK-Elk-1 signalling pathway contributing to cell survival in breast cancer. Oncogene. 2011;30(20):2367-78.
189. Gille H, Kortenjann M, Thomae O, Moomaw C, Slaughter C, Cobb MH, et al. ERK phosphorylation potentiates Elk-1-mediated ternary complex formation and transactivation. EMBO J. 1995;14(5):951-62.
190. Morris JF, Sul JY, Kim MS, Klein-Szanto AJ, Schochet T, Rustgi A, et al. Elk-1 phosphorylated at threonine-417 is present in diverse cancers and correlates with differentiation grade of colonic adenocarcinoma. Hum Pathol. 2013;44(5):766-76.
191. Rao VN, Reddy ES. elk-1 proteins interact with MAP kinases. Oncogene. 1994;9(7):1855-60.
192. Yang R, Li X, Wu Y, Zhang G, Liu X, Li Y, et al. EGFR activates GDH1 transcription to promote glutamine metabolism through MEK/ERK/ELK1 pathway in glioblastoma. Oncogene. 2020;39(14):2975-86.
193. Xiao J, Yang S, Shen P, Wang Y, Sun H, Ji F, et al. Phosphorylation of ETV4 at Ser73 by ERK kinase could block ETV4 ubiquitination degradation in colorectal cancer. Biochem Biophys Res Commun. 2017;486(4):1062-8.
194. Abe H, Okazawa M, Nakanishi S. Gene regulation via excitation and BDNF is mediated by induction and phosphorylation of the Etv1 transcription factor in cerebellar granule cells. Proc Natl Acad Sci U S A. 2012;109(22):8734-9.
195. Janknecht R. Analysis of the ERK-stimulated ETS transcription factor ER81. Mol Cell Biol. 1996;16(4):1550-6.
196. Cowley DO, Graves BJ. Phosphorylation represses Ets-1 DNA binding by reinforcing autoinhibition. Genes Dev. 2000;14(3):366-76.
197. Liu H, Grundstrom T. Calcium regulation of GM-CSF by calmodulin-dependent kinase II phosphorylation of Ets1. Mol Biol Cell. 2002;13(12):4497-507.
198. Rabault B, Ghysdael J. Calcium-induced phosphorylation of ETS1 inhibits its specific DNA binding activity. J Biol Chem. 1994;269(45):28143-51.
199. Yu JC, Chen JR, Lin CH, Zhang G, Lam PS, Wenger KH, et al. Tensile strain-induced Ets-2 phosphorylation by CaMKII and the homeostasis of cranial sutures. Plast Reconstr Surg. 2009;123(2 Suppl):83S-93S.
200. Fujiwara S, Fisher RJ, Bhat NK, Diaz de la Espina SM, Papas TS. A short-lived nuclear phosphoprotein encoded by the human ets-2 proto-oncogene is stabilized by activation of protein kinase C. Mol Cell Biol. 1988;8(11):4700-6.
201. Lindemann RK, Braig M, Ballschmieter P, Guise TA, Nordheim A, Dittmer J. Protein kinase Calpha regulates Ets1 transcriptional activity in invasive breast cancer cells. Int J Oncol. 2003;22(4):799-805.
202. Asano Y, Trojanowska M. Phosphorylation of Fli1 at threonine 312 by protein kinase C delta promotes its interaction with p300/CREB-binding protein-associated factor and subsequent acetylation in response to transforming growth factor beta. Mol Cell Biol. 2009;29(7):1882-94.
203. Bujor AM, Asano Y, Haines P, Lafyatis R, Trojanowska M. The c-Abl tyrosine kinase controls protein kinase Cdelta-induced Fli-1 phosphorylation in human dermal fibroblasts. Arthritis Rheum. 2011;63(6):1729-37.
204. Zhao H, Zhao Y, Li Z, Ouyang Q, Sun Y, Zhou D, et al. FLI1 and PKC co-activation promote highly efficient differentiation of human embryonic stem cells into endothelial-like cells. Cell Death Dis. 2018;9(2):131.
205. Baert JL, Beaudoin C, Coutte L, de Launoit Y. ERM transactivation is up-regulated by the repression of DNA binding after the PKA phosphorylation of a consensus site at the edge of the ETS domain. J Biol Chem. 2002;277(2):1002-12.
206. Wu J, Janknecht R. Regulation of the ETS transcription factor ER81 by the 90-kDa ribosomal S6 kinase 1 and protein kinase A. J Biol Chem. 2002;277(45):42669-79.
207. Lu G, Zhang Q, Huang Y, Song J, Tomaino R, Ehrenberger T, et al. Phosphorylation of ETS1 by Src family kinases prevents its recognition by the COP1 tumor suppressor. Cancer Cell. 2014;26(2):222-34.
208. Ducker C, Shaw PE. Ubiquitin-Mediated Control of ETS Transcription Factors: Roles in Cancer and Development. Int J Mol Sci. 2021;22(10).
209. Rieske P, Pongubala JM. AKT induces transcriptional activity of PU.1 through phosphorylation-mediated modifications within its transactivation domain. J Biol Chem. 2001;276(11):8460-8.
210. Seshire A, Rossiger T, Frech M, Beez S, Hagemeyer H, Puccetti E. Direct interaction of PU.1 with oncogenic transcription factors reduces its serine phosphorylation and promoter binding. Leukemia. 2012;26(6):1338-47.
211. Ray-Gallet D, Moreau-Gachelin F. Phosphorylation of the Spi-B transcription factor reduces its intrinsic stability. FEBS Lett. 1999;464(3):164-8.
212. Di Cerbo V, Schneider R. Cancers with wrong HATs: the impact of acetylation. Brief Funct Genomics. 2013;12(3):231-43.
213. Di Martile M, Del Bufalo D, Trisciuoglio D. The multifaceted role of lysine acetylation in cancer: prognostic biomarker and therapeutic target. Oncotarget. 2016;7(34):55789-810.
214. Park JM, Jo SH, Kim MY, Kim TH, Ahn YH. Role of transcription factor acetylation in the regulation of metabolic homeostasis. Protein Cell. 2015;6(11):804-13.
215. Thiagarajan D, Vedantham S, Ananthakrishnan R, Schmidt AM, Ramasamy R. Mechanisms of transcription factor acetylation and consequences in hearts. Biochim Biophys Acta. 2016;1862(12):2221-31.
216. Dong Z. Acetylation of Ets-1 is the key to chromatin remodeling for miR-192 expression. Sci Signal. 2013;6(278):pe21.
217. Kato M, Dang V, Wang M, Park JT, Deshpande S, Kadam S, et al. TGF-beta induces acetylation of chromatin and of Ets-1 to alleviate repression of miR-192 in diabetic nephropathy. Sci Signal. 2013;6(278):ra43.
218. Chen J, Fu Y, Day DS, Sun Y, Wang S, Liang X, et al. VEGF amplifies transcription through ETS1 acetylation to enable angiogenesis. Nat Commun. 2017;8(1):383.
219. Blee AM, Liu S, Wang L, Huang H. BET bromodomain-mediated interaction between ERG and BRD4 promotes prostate cancer cell invasion. Oncotarget. 2016;7(25):38319-32.
220. Roe JS, Mercan F, Rivera K, Pappin DJ, Vakoc CR. BET Bromodomain Inhibition Suppresses the Function of Hematopoietic Transcription Factors in Acute Myeloid Leukemia. Mol Cell. 2015;58(6):1028-39.
221. Asano Y, Czuwara J, Trojanowska M. Transforming growth factor-beta regulates DNA binding activity of transcription factor Fli1 by p300/CREB-binding protein-associated factor-dependent acetylation. J Biol Chem. 2007;282(48):34672-83.
222. Asano Y, Trojanowska M. Fli1 represses transcription of the human alpha2(I) collagen gene by recruitment of the HDAC1/p300 complex. PLoS One. 2013;8(9):e74930.
223. Schlottmann S, Erkizan HV, Barber-Rotenberg JS, Knights C, Cheema A, Uren A, et al. Acetylation Increases EWS-FLI1 DNA Binding and Transcriptional Activity. Front Oncol. 2012;2:107.
224. Goel A, Janknecht R. Acetylation-mediated transcriptional activation of the ETS protein ER81 by p300, P/CAF, and HER2/Neu. Mol Cell Biol. 2003;23(17):6243-54.
225. Goel A, Janknecht R. Concerted activation of ETS protein ER81 by p160 coactivators, the acetyltransferase p300 and the receptor tyrosine kinase HER2/Neu. J Biol Chem. 2004;279(15):14909-16.
226. Guo B, Panagiotaki N, Warwood S, Sharrocks AD. Dynamic modification of the ETS transcription factor PEA3 by sumoylation and p300-mediated acetylation. Nucleic Acids Res. 2011;39(15):6403-13.
227. Jiang W, Xu Y, Chen X, Pan S, Zhu X. E26 transformation-specific variant 4 as a tumor promotor in human cancers through specific molecular mechanisms. Mol Ther Oncolytics. 2021;22:518-27.
228. Putnik J, Zhang CD, Archangelo LF, Tizazu B, Bartels S, Kickstein M, et al. The interaction of ETV6 (TEL) and TIP60 requires a functional histone acetyltransferase domain in TIP60. Biochim Biophys Acta. 2007;1772(11-12):1211-24.
229. Li X, Li S, Li B, Li Y, Aman S, Xia K, et al. Acetylation of ELF5 suppresses breast cancer progression by promoting its degradation and targeting CCND1. NPJ Precis Oncol. 2021;5(1):20.
230. Ohtake F, Saeki Y, Ishido S, Kanno J, Tanaka K. The K48-K63 Branched Ubiquitin Chain Regulates NF-kappaB Signaling. Mol Cell. 2016;64(2):251-66.
231. Ohtake F, Tsuchiya H, Saeki Y, Tanaka K. K63 ubiquitylation triggers proteasomal degradation by seeding branched ubiquitin chains. Proc Natl Acad Sci U S A. 2018;115(7):E1401-E8.
232. Wang G, Gao Y, Li L, Jin G, Cai Z, Chao JI, et al. K63-linked ubiquitination in kinase activation and cancer. Front Oncol. 2012;2:5.
233. Ji Z, Degerny C, Vintonenko N, Deheuninck J, Foveau B, Leroy C, et al. Regulation of the Ets-1 transcription factor by sumoylation and ubiquitinylation. Oncogene. 2007;26(3):395-406.
234. Baert JL, Monte D, Verreman K, Degerny C, Coutte L, de Launoit Y. The E3 ubiquitin ligase complex component COP1 regulates PEA3 group member stability and transcriptional activity. Oncogene. 2010;29(12):1810-20.
235. Vitari AC, Leong KG, Newton K, Yee C, O'Rourke K, Liu J, et al. COP1 is a tumour suppressor that causes degradation of ETS transcription factors. Nature. 2011;474(7351):403-6.
236. Migliorini D, Bogaerts S, Defever D, Vyas R, Denecker G, Radaelli E, et al. Cop1 constitutively regulates c-Jun protein stability and functions as a tumor suppressor in mice. J Clin Invest. 2011;121(4):1329-43.
237. Ouyang M, Wang H, Ma J, Lu W, Li J, Yao C, et al. COP1, the negative regulator of ETV1, influences prognosis in triple-negative breast cancer. BMC Cancer. 2015;15:132.
238. Sawada G, Ueo H, Matsumura T, Uchi R, Ishibashi M, Mima K, et al. Loss of COP1 expression determines poor prognosisin patients with gastric cancer. Oncol Rep. 2013;30(4):1971-5.
239. Song Y, Liu Y, Pan S, Xie S, Wang ZW, Zhu X. Role of the COP1 protein in cancer development and therapy. Semin Cancer Biol. 2020;67(Pt 2):43-52.
240. Gan W, Dai X, Lunardi A, Li Z, Inuzuka H, Liu P, et al. SPOP Promotes Ubiquitination and Degradation of the ERG Oncoprotein to Suppress Prostate Cancer Progression. Mol Cell. 2015;59(6):917-30.
241. Shoag J, Liu D, Blattner M, Sboner A, Park K, Deonarine L, et al. SPOP mutation drives prostate neoplasia without stabilizing oncogenic transcription factor ERG. J Clin Invest. 2018;128(1):381-6.
242. Hong Z, Zhang W, Ding D, Huang Z, Yan Y, Cao W, et al. DNA Damage Promotes TMPRSS2-ERG Oncoprotein Destruction and Prostate Cancer Suppression via Signaling Converged by GSK3beta and WEE1. Mol Cell. 2020;79(6):1008-23 e4.
243. Yeh CH, Bellon M, Nicot C. FBXW7: a critical tumor suppressor of human cancers. Mol Cancer. 2018;17(1):115.
244. Gierisch ME, Pfistner F, Lopez-Garcia LA, Harder L, Schafer BW, Niggli FK. Proteasomal Degradation of the EWS-FLI1 Fusion Protein Is Regulated by a Single Lysine Residue. J Biol Chem. 2016;291(52):26922-33.
245. Su S, Chen J, Jiang Y, Wang Y, Vital T, Zhang J, et al. SPOP and OTUD7A Control EWS-FLI1 Protein Stability to Govern Ewing Sarcoma Growth. Adv Sci (Weinh). 2021;8(14):e2004846.
246. Molina MD, Quirin M, Haillot E, De Croze N, Range R, Rouel M, et al. MAPK and GSK3/ss-TRCP-mediated degradation of the maternal Ets domain transcriptional repressor Yan/Tel controls the spatial expression of nodal in the sea urchin embryo. PLoS Genet. 2018;14(9):e1007621.
247. Roukens MG, Alloul-Ramdhani M, Moghadasi S, Op den Brouw M, Baker DA. Downregulation of vertebrate Tel (ETV6) and Drosophila Yan is facilitated by an evolutionarily conserved mechanism of F-box-mediated ubiquitination. Mol Cell Biol. 2008;28(13):4394-406.
248. Quintero-Barceinas RS, Gehringer F, Ducker C, Saxton J, Shaw PE. ELK-1 ubiquitination status and transcriptional activity are modulated independently of F-Box protein FBXO25. J Biol Chem. 2021;296:100214.
249. Ducker C, Chow LKY, Saxton J, Handwerger J, McGregor A, Strahl T, et al. De-ubiquitination of ELK-1 by USP17 potentiates mitogenic gene expression and cell proliferation. Nucleic Acids Res. 2019;47(9):4495-508.
250. Evans EL, Saxton J, Shelton SJ, Begitt A, Holliday ND, Hipskind RA, et al. Dimer formation and conformational flexibility ensure cytoplasmic stability and nuclear accumulation of Elk-1. Nucleic Acids Res. 2011;39(15):6390-402.
251. Teixeira FR, Manfiolli AO, Soares CS, Baqui MM, Koide T, Gomes MD. The F-box protein FBXO25 promotes the proteasome-dependent degradation of ELK-1 protein. J Biol Chem. 2013;288(39):28152-62.
252. Chen X, Qin Y, Zhang Z, Xing Z, Wang Q, Lu W, et al. Hyper-SUMOylation of ERG Is Essential for the Progression of Acute Myeloid Leukemia. Front Mol Biosci. 2021;8:652284.
253. van den Akker E, Ano S, Shih HM, Wang LC, Pironin M, Palvimo JJ, et al. FLI-1 functionally interacts with PIASxalpha, a member of the PIAS E3 SUMO ligase family. J Biol Chem. 2005;280(45):38035-46.
254. Bojovic BB, Hassell JA. The transactivation function of the Pea3 subfamily Ets transcription factors is regulated by sumoylation. DNA Cell Biol. 2008;27(6):289-305.
255. Oh S, Shin S, Janknecht R. Sumoylation of transcription factor ETV1 modulates its oncogenic potential in prostate cancer. Int J Clin Exp Pathol. 2021;14(7):795-810.
256. Roukens MG, Alloul-Ramdhani M, Vertegaal AC, Anvarian Z, Balog CI, Deelder AM, et al. Identification of a new site of sumoylation on Tel (ETV6) uncovers a PIAS-dependent mode of regulating Tel function. Mol Cell Biol. 2008;28(7):2342-57.
257. Chakrabarti SR, Sood R, Nandi S, Nucifora G. Posttranslational modification of TEL and TEL/AML1 by SUMO-1 and cell-cycle-dependent assembly into nuclear bodies. Proc Natl Acad Sci U S A. 2000;97(24):13281-5.
258. Chowdhury D, Singh A, Gupta A, Tulsawani R, Meena RC, Chakrabarti A. p38 MAPK pathway-dependent SUMOylation of Elk-1 and phosphorylation of PIAS2 correlate with the downregulation of Elk-1 activity in heat-stressed HeLa cells. Cell Stress Chaperones. 2019;24(2):393-407.
259. Deng L, Meng T, Chen L, Wei W, Wang P. The role of ubiquitination in tumorigenesis and targeted drug discovery. Signal Transduct Target Ther. 2020;5(1):11.
260. Rosonina E, Akhter A, Dou Y, Babu J, Sri Theivakadadcham VS. Regulation of transcription factors by sumoylation. Transcription. 2017;8(4):220-31.
261. Zhao Q, Ma Y, Li Z, Zhang K, Zheng M, Zhang S. The Function of SUMOylation and Its Role in the Development of Cancer Cells under Stress Conditions: A Systematic Review. Stem Cells Int. 2020;2020:8835714.
262. Zoma M, Curti L, Shinde D, Albino D, Mitra A, Sgrignani J, et al. EZH2-induced lysine K362 methylation enhances TMPRSS2-ERG oncogenic activity in prostate cancer. Nat Commun. 2021;12(1):4147.
263. Tsokos GC, Nambiar MP, Juang YT. Activation of the Ets transcription factor Elf-1 requires phosphorylation and glycosylation: defective expression of activated Elf-1 is involved in the decreased TCR zeta chain gene expression in patients with systemic lupus erythematosus. Ann N Y Acad Sci. 2003;987:240-5.
264. Hsing M, Wang Y, Rennie PS, Cox ME, Cherkasov A. ETS transcription factors as emerging drug targets in cancer. Med Res Rev. 2020;40(1):413-30.
265. Lambert M, Jambon S, Depauw S, David-Cordonnier MH. Targeting Transcription Factors for Cancer Treatment. Molecules. 2018;23(6).
266. Maroulakou IG, Bowe DB. Expression and function of Ets transcription factors in mammalian development: a regulatory network. Oncogene. 2000;19(55):6432-42.
267. Oliver JR, Kushwah R, Hu J. Multiple roles of the epithelium-specific ETS transcription factor, ESE-1, in development and disease. Lab Invest. 2012;92(3):320-30.
268. Suico MA, Shuto T, Kai H. Roles and regulations of the ETS transcription factor ELF4/MEF. J Mol Cell Biol. 2017;9(3):168-77.
269. Feng FY, Brenner JC, Hussain M, Chinnaiyan AM. Molecular pathways: targeting ETS gene fusions in cancer. Clin Cancer Res. 2014;20(17):4442-8.
270. Kumar-Sinha C, Kalyana-Sundaram S, Chinnaiyan AM. Landscape of gene fusions in epithelial cancers: seq and ye shall find. Genome Med. 2015;7:129.
271. Prasad P, Roy SS. Glutamine regulates ovarian cancer cell migration and invasion through ETS1. Heliyon. 2021;7(5):e07064.
272. Smith AM, Findlay VJ, Bandurraga SG, Kistner-Griffin E, Spruill LS, Liu A, et al. ETS1 transcriptional activity is increased in advanced prostate cancer and promotes the castrate-resistant phenotype. Carcinogenesis. 2012;33(3):572-80.
273. Dhara A, Aier I, Paladhi A, Varadwaj PK, Hira SK, Sen N. PGC1 alpha coactivates ERG fusion to drive antioxidant target genes under metabolic stress. Commun Biol. 2022;5(1):416.
274. Nodehi RS, Kalantari B, Raafat J, Ansarinejad N, Moazed V, Mortazavizadeh SMR, et al. A randomized, double-blind, phase III, non-inferiority clinical trial comparing the efficacy and safety of TA4415V (a proposed Trastuzumab biosimilar) and Herceptin (Trastuzumab reference product) in HER2-positive early-stage breast Cancer patients. BMC Pharmacology and Toxicology. 2022;23(1):57.
275. Sizemore GM, Pitarresi JR, Balakrishnan S, Ostrowski MC. The ETS family of oncogenic transcription factors in solid tumours. Nature Reviews Cancer. 2017;17(6):337-51.
276. Spriano F, Chung EYL, Gaudio E, Tarantelli C, Cascione L, Napoli S, et al. The ETS Inhibitors YK-4-279 and TK-216 Are Novel Antilymphoma Agents. Clin Cancer Res. 2019;25(16):5167-76.
277. Taniguchi T, Iizumi Y, Watanabe M, Masuda M, Morita M, Aono Y, et al. Resveratrol directly targets DDX5 resulting in suppression of the mTORC1 pathway in prostate cancer. Cell Death Dis. 2016;7:e2211.
278. Shao L, Kahraman N, Yan G, Wang J, Ozpolat B, Ittmann M. Targeting the TMPRSS2/ERG fusion mRNA using liposomal nanovectors enhances docetaxel treatment in prostate cancer. Prostate. 2020;80(1):65-73.
279. Shao L, Tekedereli I, Wang J, Yuca E, Tsang S, Sood A, et al. Highly specific targeting of the TMPRSS2/ERG fusion gene using liposomal nanovectors. Clin Cancer Res. 2012;18(24):6648-57.
280. Urbinati G, Ali HM, Rousseau Q, Chapuis H, Desmaele D, Couvreur P, et al. Antineoplastic Effects of siRNA against TMPRSS2-ERG Junction Oncogene in Prostate Cancer. PLoS One. 2015;10(5):e0125277.
281. Magistroni V, Mologni L, Sanselicio S, Reid JF, Redaelli S, Piazza R, et al. ERG deregulation induces PIM1 over-expression and aneuploidy in prostate epithelial cells. PLoS One. 2011;6(11):e28162.
282. Mortensen L, Ordulu Z, Dagogo-Jack I, Bossuyt V, Winters L, Taghian A, et al. Locally Recurrent Secretory Carcinoma of the Breast with NTRK3 Gene Fusion. Oncologist. 2021;26(10):818-24.
283. Kaindl U, Morak M, Portsmouth C, Mecklenbrauker A, Kauer M, Zeginigg M, et al. Blocking ETV6/RUNX1-induced MDM2 overexpression by Nutlin-3 reactivates p53 signaling in childhood leukemia. Leukemia. 2014;28(3):600-8.
284. Kayarthodi S, Fujimura Y, Fang J, Morsalin S, Rao VN, Reddy ES. Anti-Epileptic Drug Targets Ewing Sarcoma. J Pharm Sci Pharmacol. 2014;1(2):87-100.