Abstract
This commentary contextualizes the findings reported in Soares et al. (Scientific Reports, 2025), which investigated maternal–newborn humoral immune interactions during Zika virus (ZIKV) infection using epitope-level mapping. In that study, paired sera from mother–newborn dyads were analyzed using SPOT-synthesis peptide arrays spanning the ZIKV proteome, revealing a high qualitative overlap of antibody targets between mothers and neonates, alongside marked quantitative differences in signal intensity. Immunodominant responses were primarily directed against Envelope (E) and NS5 proteins, while Capsid (C) and NS1 displayed differential recognition patterns. Here, we discuss the biological significance of these observations, emphasizing how transplacental IgG transfer preserves antigenic targeting while modulating response magnitude. We further explore translational implications for flavivirus sero-diagnostics and maternal vaccination strategies. No new experimental data are presented in this commentary.
Keywords
Zika virus, Congenital Zika, Maternal immunity, Neonatal immunity, Antibody transfer, B-cell epitopes, Flavivirus diagnostics
Introduction
The Zika virus (ZIKV) epidemic in Brazil during 2015–2016 reshaped global understanding of arboviral disease by revealing the virus’s capacity for vertical transmission and its association with congenital abnormalities. While extensive work focused on viral neurotropism and fetal pathology, the immunological interface between infected mothers and their newborns has remained comparatively underexplored.
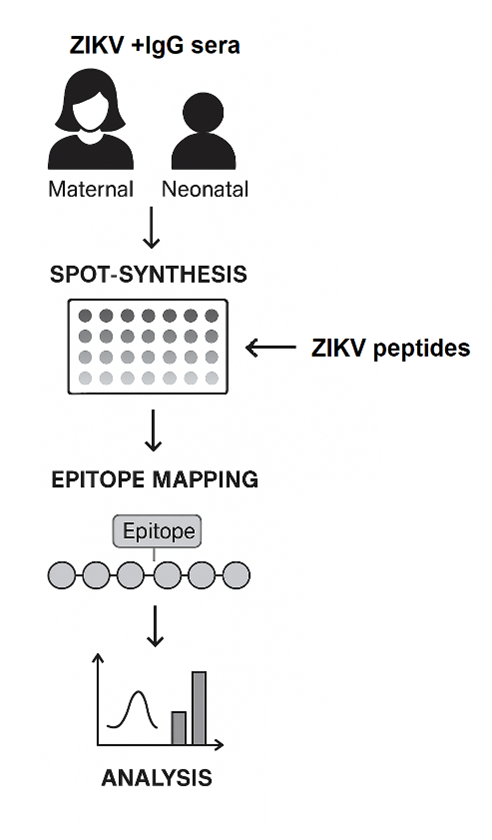
This commentary builds upon our recent research [1], in which paired sera from mother–newborn dyads collected at birth (cord blood) were analyzed to characterize the fine specificity of anti-ZIKV IgG responses. That study included 2 mother–newborn pairs, all IgG-positive for ZIKV, and focused on linear B-cell epitope recognition across the viral proteome. Here, we synthesize those findings and discuss their broader biological and translational implications, without presenting new experimental data.
Intergenerational Epitope Landscapes: What the Data Reveal
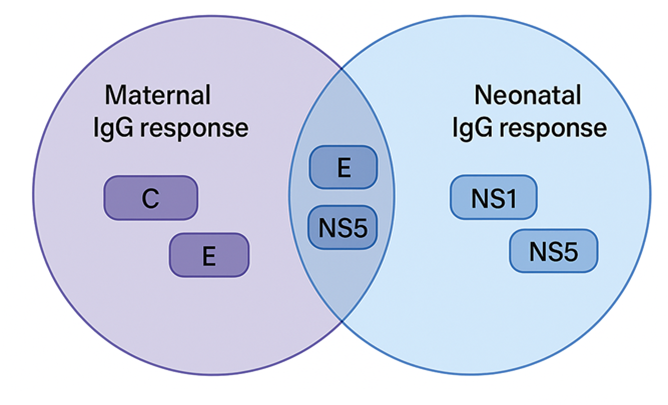
Using SPOT-synthesis peptide arrays comprising 719 overlapping peptides covering the full ZIKV polyprotein, the original study identified 148 ZIKV-specific linear B-cell epitopes after exclusion of cross-reactive regions recognized by dengue virus (DENV)-positive controls. Maternal and neonatal sera exhibited substantial qualitative overlap in epitope recognition (approximately 41–52%), particularly within the Envelope (E) and NS5 proteins.
Despite this shared antigenic landscape, quantitative differences in antibody binding intensity were consistently observed between mothers and newborns. These findings suggest that while placental IgG transfer conserves which viral regions are recognized, it modulates the strength of their targeting. In simpler terms, antibodies cross the placenta and retain the same targets, but the strength of the response changes in the newborn.
Biological Interpretation: Immune Inheritance Across the Placenta
Placental transfer of maternal IgG, primarily mediated by the neonatal Fc receptor (FcRn) [2], is most efficient during late gestation and results in neonatal antibody repertoires that closely resemble those of the mother. However, the epitope-level differences observed in the original study indicate that neonatal immunity is not merely a passive mirror of maternal responses.
Selective transfer, differential decay rates after birth, and potential in utero antigen exposure may all contribute to shaping distinct postnatal antibody landscapes. These processes support the concept of immune imprinting, whereby maternal infection leaves molecular signatures that persist into early life and influence subsequent immune responsiveness [3–5].
Methodological Perspective: Why Epitope-Level Mapping Matters
Conventional serological assays measure antibody responses at the protein or whole-virus level, often obscuring fine specificity and cross-reactivity. Epitope-level mapping using SPOT-synthesis enables amino acid–resolution analysis of humoral immunity, allowing precise discrimination between ZIKV-specific and cross-reactive flavivirus responses [6–8].
In the context of pregnancy, this resolution is particularly valuable, as it reveals how immune information is transmitted across generations at a molecular scale. By integrating experimental peptide arrays with structural predictions, the original study provided mechanistic insight into why certain viral regions dominate maternal and neonatal antibody recognition.
Translational Outlook and Conceptual Framework
The identification of ZIKV-specific epitopes has clear implications for improving serological diagnostics in regions where multiple flaviviruses co-circulate. Epitope-informed assays may reduce false positives arising from cross-reactive antibodies, a persistent challenge in flavivirus-endemic areas.
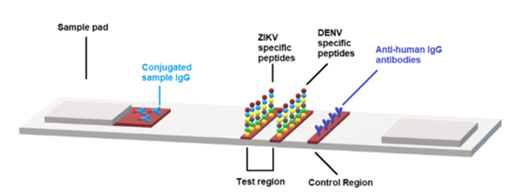
In addition, dominant and efficiently transferred epitopes, particularly within E and NS5, represent promising targets for maternal immunization strategies aimed at maximizing neonatal protection [2,9,10]. Figure 3 illustrates a conceptual peptide-based lateral flow immunoassay, intended solely as a future diagnostic framework; this assay has not yet been experimentally validated.
Concluding Perspective
The findings discussed here support the view that vertical immunity is an active and selective biological process rather than a simple transfer of maternal antibodies. Epitope-level mapping reveals how immune recognition patterns are conserved yet reshaped across the mother–newborn interface, offering new insight into immune inheritance during congenital viral infections.
By clarifying how maternal ZIKV infection imprints neonatal antibody landscapes, this work contributes to a broader understanding of developmental immunology. It provides a conceptual foundation for improved diagnostics and preventive strategies against flaviviruses.
Conflict of Interest
The authors declare no competing interests.
Acknowledgments
We thank the participating mothers and infants, as well as our collaborators at UFMG and FMJ. This work was supported by FAPESP, CNPq, and Fiocruz/Pasteur/USP.
References
2. Hanson LA, Korotkova M, Lundin S, Håversen L, Silfverdal SA, Mattsby-Baltzer I, et al. The transfer of immunity from mother to child. Ann N Y Acad Sci. 2003 Apr;987:199–206.
3. Robbiani DF, Bozzacco L, Keeffe JR, Khouri R, Olsen PC, Gazumyan A, et al. Recurrent Potent Human Neutralizing Antibodies to Zika Virus in Brazil and Mexico. Cell. 2017 May 4;169(4):597–609.e11.
4. Zhao H, Fernandez E, Dowd KA, Speer SD, Platt DJ, Gorman MJ, et al. Structural Basis of Zika Virus-Specific Antibody Protection. Cell. 2016 Aug 11;166(4):1016–27.
5. Palmeira P, Quinello C, Silveira-Lessa AL, Zago CA, Carneiro-Sampaio M. IgG placental transfer in healthy and pathological pregnancies. Clin Dev Immunol. 2012;2012:985646.
6. Frank R. The SPOT-synthesis technique. Synthetic peptide arrays on membrane supports--principles and applications. J Immunol Methods. 2002 Sep 1;267(1):13–26.
7. Volkmer R. Synthesis and application of peptide arrays: quo vadis SPOT technology. Chembiochem. 2009 Jun 15;10(9):1431–42.
8. Freire MCLC, Pol-Fachin L, Coêlho DF, Viana IFT, Magalhães T, Cordeiro MT, et al. Mapping Putative B-Cell Zika Virus NS1 Epitopes Provides Molecular Basis for Anti-NS1 Antibody Discrimination between Zika and Dengue Viruses. ACS Omega. 2017 Jul 31;2(7):3913–20.
9. Sirohi D, Chen Z, Sun L, Klose T, Pierson TC, Rossmann MG, et al. The 3.8 Å resolution cryo-EM structure of Zika virus. Science. 2016 Apr 22;352(6284):467–70.
10. Reyes-Sandoval A, Ludert JE. The Dual Role of the Antibody Response Against the Flavivirus Non-structural Protein 1 (NS1) in Protection and Immuno-Pathogenesis. Front Immunol. 2019 Jul 18;10:1651.